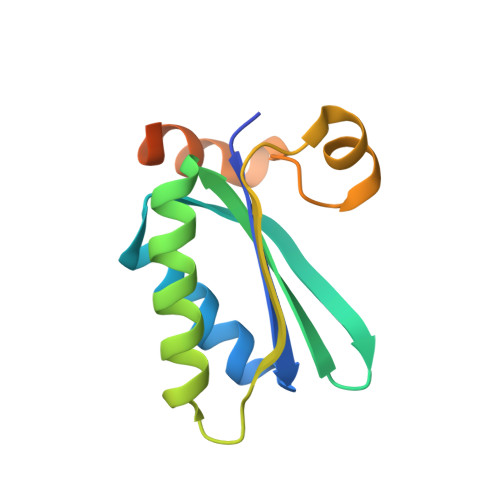
Protein structures forming the shell of primitive bacterial organelles
Kerfeld, C.A., Sawaya, M.R., Tanaka, S., Nguyen, C.V., Phillips, M., Beeby, M., Yeates, T.O.(2005) Science 309: 936-938
- PubMed: 16081736 Search on PubMed
- DOI: https://doi.org/10.1126/science.1113397
- Primary Citation Related Structures:
2A10, 2A18, 2A1B - PubMed Abstract:
Bacterial microcompartments are primitive organelles composed entirely of protein subunits. Genomic sequence databases reveal the widespread occurrence of microcompartments across diverse microbes. The prototypical bacterial microcompartment is the carboxysome, a protein shell for sequestering carbon fixation reactions. We report three-dimensional crystal structures of multiple carboxysome shell proteins, revealing a hexameric unit as the basic microcompartment building block and showing how these hexamers assemble to form flat facets of the polyhedral shell. The structures suggest how molecular transport across the shell may be controlled and how structural variations might govern the assembly and architecture of these subcellular compartments.
- Molecular Biology Institute, University of California, Los Angeles (UCLA), Box 951570, Los Angeles, CA 90095, USA.
Organizational Affiliation: