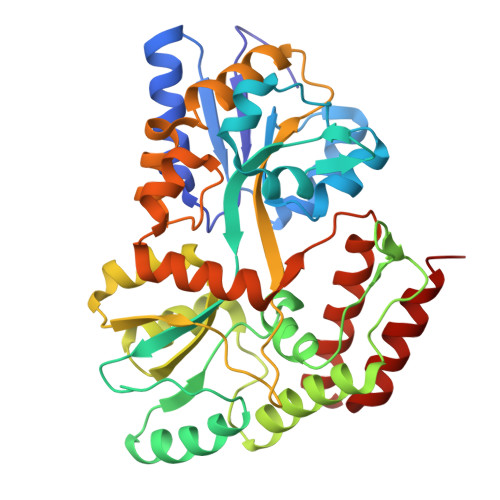
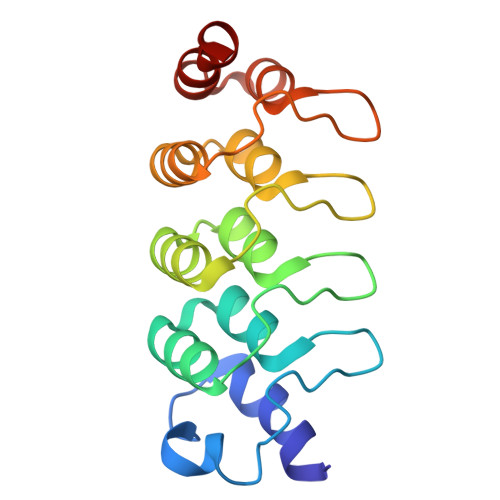
An engineered symmetrical scaffold system enables high resolution imaging of small cargo proteins by cryo-electron microscopy
Ferreira, D.S.M., Noble, M., Rowland, R.J., Gittins, O., Martin, M., Endicott, J., Fairhead, M., von Delft, F., Pike, A.C.W., Sauer, D.B., Dlamini, L.S.To be published.