Yeast Display Technology Enables Rapid Discovery of Low-Nanomolar Macrocyclic Peptide Inhibitors of Human Angiotensin-Converting Enzyme 2.
Romanyuk, Z., Bettin, G., Brear, P., Linciano, S., Mazzocato, Y., Bonadies, S., Zanotto, I., Mazzucco, C., Monferone, A., Soler, M.A., Pasut, G., Martin, S., Scarso, A., Heinis, C., Rothenberger, S., Hyvonen, M., Angelini, A.(2026) J Med Chem
- PubMed: 41875055
- DOI: https://doi.org/10.1021/acs.jmedchem.5c02876
- Primary Citation Related Structures:
28KD, 9RVT, 9SPA - PubMed Abstract:
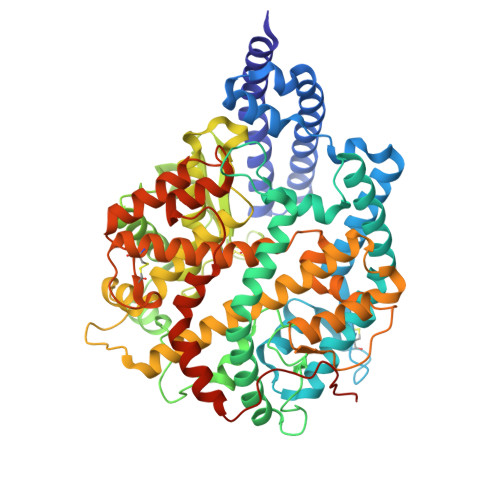
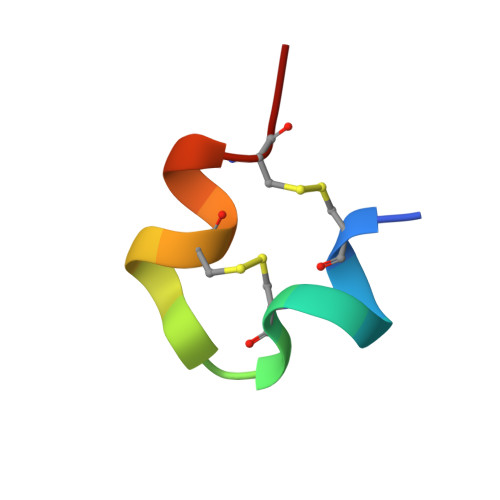
Macrocyclic peptides (MPs) are valuable molecular formats for drug development, bridging small molecules and larger biologics due to their favorable pharmacological properties. Here, we describe the discovery of low-nanomolar inhibitors of human angiotensin-converting enzyme 2 (hACE2) by quantitatively screening millions of structurally diverse disulfide-cyclized peptide ligands using yeast display technology. The most potent yeast-encoded "one-ring" and "two-ring" MP inhibit hACE2 with K i values of 1.9 and 1.5 nM, respectively. These inhibitory potencies are comparable to those of other cyclic peptides discovered using well-established in vitro display technologies. Crystal structures of the two MPs in complex with hACE2 reveal the adoption of either a rigid β-hairpin or a cysteine-stabilized α-helix/α-helix motif. Both MPs exhibit binding modes distinct from those of previously reported inhibitors. Thus, yeast display is a valid technology to rapidly generate MPs with desired binding properties for the development of potential therapeutics.
- Department of Molecular Sciences and Nanosystems, Ca' Foscari University of Venice, Via Torino 155, 30172 Mestre, Italy.
Organizational Affiliation: