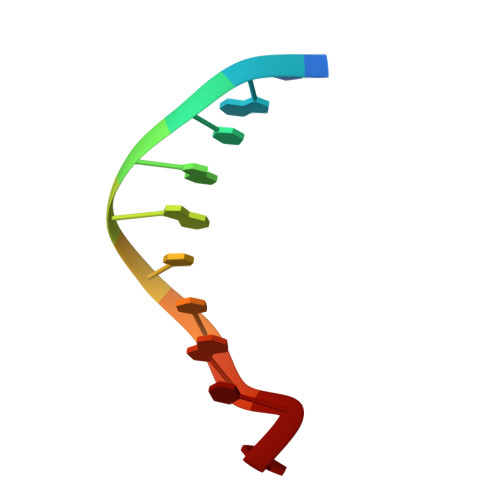
Crystal structure of a DNA decamer showing a novel pseudo four-way helix-helix junction.
Spink, N., Nunn, C.M., Vojtechovsky, J., Berman, H.M., Neidle, S.(1995) Proc Natl Acad Sci U S A 92: 10767-10771
- PubMed: 7479880 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.92.23.10767
- Primary Citation Related Structures:
237D - PubMed Abstract:
The crystal structure of the decanucleotide d(CGCAATTGCG)2 has been solved by a combination of molecular replacement and heavy-atom procedures and has been refined to an R factor of 20.2% at 2.7 A. It is not a fully base-paired duplex but has a central core of eight Watson-Crick base pairs flanked by unpaired terminal guanosines and cytosines. These participate in hydrogen-bonding arrangements with adjacent decamer duplexes in the crystal lattice. The unpaired guanosines are bound in the G+C regions of duplex minor grooves. The cytosines have relatively high mobility, even though they are constrained to be in one region where they are involved in base-paired triplets with G.C base pairs. The 5'-AATT sequence in the duplex region has a narrow minor groove, providing further confirmation of the sequence-dependent nature of groove width.
- Cancer Research Campaign Biomolecular Structure Unit, Institute of Cancer Research, Sutton, Surrey, United Kingdom.
Organizational Affiliation: