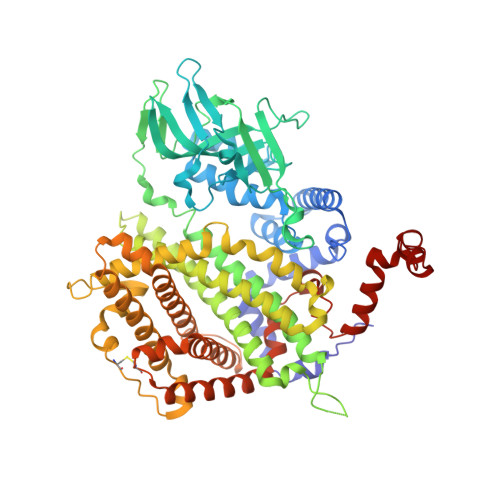
Acyl-Coa Oxidase 1 from Arabidopsis Thaliana. Structure of a Key Enzyme in Plant Lipid Metabolism
Pedersen, L., Henriksen, A.(2005) J Mol Biology 345: 487
- PubMed: 15581893 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.10.062
- Primary Citation Related Structures:
1W07 - PubMed Abstract:
The peroxisomal acyl-CoA oxidase family plays an essential role in lipid metabolism by catalyzing the conversion of acyl-CoA into trans-2-enoyl-CoA during fatty acid beta-oxidation. Here, we report the X-ray structure of the FAD-containing Arabidopsis thaliana acyl-CoA oxidase 1 (ACX1), the first three-dimensional structure of a plant acyl-CoA oxidase. Like other acyl-CoA oxidases, the enzyme is a dimer and it has a fold resembling that of mammalian acyl-CoA oxidase. A comparative analysis including mammalian acyl-CoA oxidase and the related tetrameric mitochondrial acyl-CoA dehydrogenases reveals a substrate-binding architecture that explains the observed preference for long-chained, mono-unsaturated substrates in ACX1. Two anions are found at the ACX1 dimer interface and for the first time the presence of a disulfide bridge in a peroxisomal protein has been observed. The functional differences between the peroxisomal acyl-CoA oxidases and the mitochondrial acyl-CoA dehydrogenases are attributed to structural differences in the FAD environments.
- Carlsberg Laboratory, Gamle Carlsberg Vej 10, DK-2500 Valby, Denmark.
Organizational Affiliation: