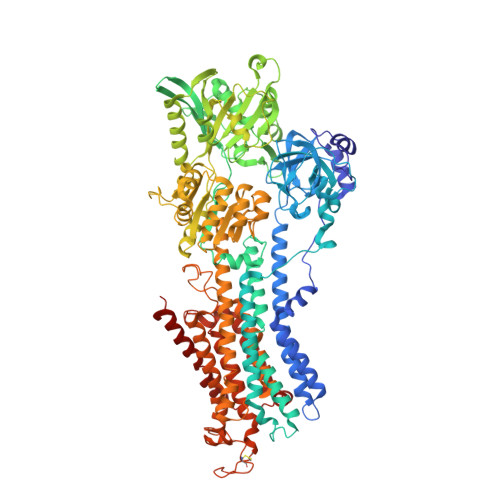
Crystal structure of the calcium pump with a bound ATP analogue.
Toyoshima, C., Mizutani, T.(2004) Nature 430: 529-535
- PubMed: 15229613 Search on PubMed
- DOI: https://doi.org/10.1038/nature02680
- Primary Citation Related Structures:
1VFP - PubMed Abstract:
P-type ATPases are ATP-powered ion pumps that establish ion concentration gradients across cell and organelle membranes. Here, we describe the crystal structure of the Ca2+ pump of skeletal muscle sarcoplasmic reticulum, a representative member of the P-type ATPase superfamily, with an ATP analogue, a Mg2+ and two Ca2+ ions in the respective binding sites. In this state, the ATP analogue reorganizes the three cytoplasmic domains (A, N and P), which are widely separated without nucleotide, by directly bridging the N and P domains. The structure of the P-domain itself is altered by the binding of the ATP analogue and Mg2+. As a result, the A-domain is tilted so that one of the transmembrane helices moves to lock the cytoplasmic gate of the transmembrane Ca2+-binding sites. This appears to be the mechanism for occluding the bound Ca2+ ions, before releasing them into the lumen of the sarcoplasmic reticulum.
- Institute of Molecular and Cellular Biosciences, The University of Tokyo, Bunkyo-ku, Tokyo 113-0032, Japan. ct@iam.u-tokyo.ac.jp
Organizational Affiliation: