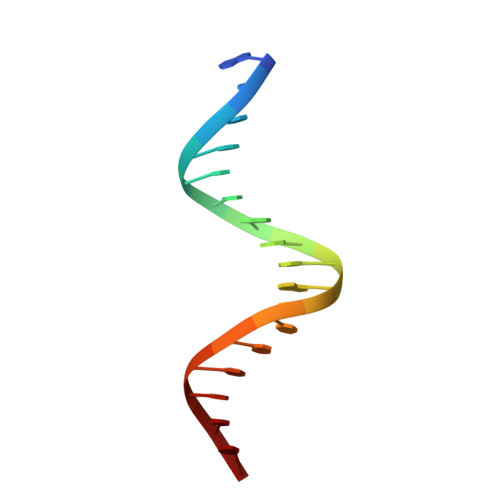
Pre-organized structure of viral DNA at the binding-processing site of HIV-1 integrase
Renisio, J.G., Cosquer, S., Cherrak, I., El Antri, S., Mauffret, O., Fermandjian, S.(2005) Nucleic Acids Res 33: 1970-1981
- PubMed: 15814814 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gki346
- Primary Citation Related Structures:
1TQR - PubMed Abstract:
The integration of the human immunodeficiency virus type 1 DNA into the host cell genome is catalysed by the viral integrase (IN). The reaction consists of a 3'-processing [dinucleotide released from each 3' end of the viral long terminal repeat (LTR)] followed by a strand transfer (insertion of the viral genome into the human chromosome). A 17 base pair oligonucleotide d(GGAAAATCTCTAGCAGT), d(ACTGCTAGAGATTTTCC) reproducing the U5-LTR extremity of viral DNA that contains the IN attachment site was analysed by NMR using the classical NOEs and scalar coupling constants in conjunction with a small set of residual dipolar coupling constants (RDCs) measured at the 13C/15N natural abundance. The combination of these two types of parameters in calculations significantly improved the DNA structure determination. The well-known features of A-tracts were clearly identified by RDCs in the first part of the molecule. The binding/cleavage site at the viral DNA end is distinguishable by a loss of regular base stacking and a distorted minor groove that can aid its specific recognition by IN.
- Centre National de la Recherche Scientifique Unité Mixte de Recherche 8113, Laboratoire de Biotechnologies et Pharmacologie génétique Appliquée, Ecole Normale Supérieure de Cachan 94235 Cachan, France.
Organizational Affiliation: