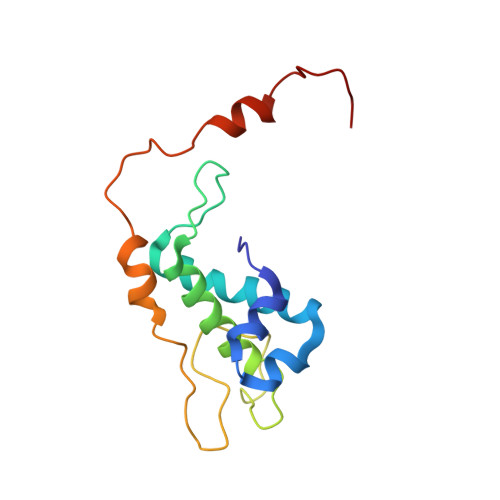
NMR structure of protein yqbG from Bacillus subtilis reveals a novel alpha-helical protein fold.
Liu, G., Shen, Y., Xiao, R., Acton, T., Ma, L.C., Joachimiak, A., Montelione, G.T., Szyperski, T.(2006) Proteins 62: 288-291
- PubMed: 16281282 Search on PubMed
- DOI: https://doi.org/10.1002/prot.20666
- Primary Citation Related Structures:
1ZTS - Department of Chemistry, University at Buffalo, The State University of New York, Buffalo, New York 14260, USA.
Organizational Affiliation: