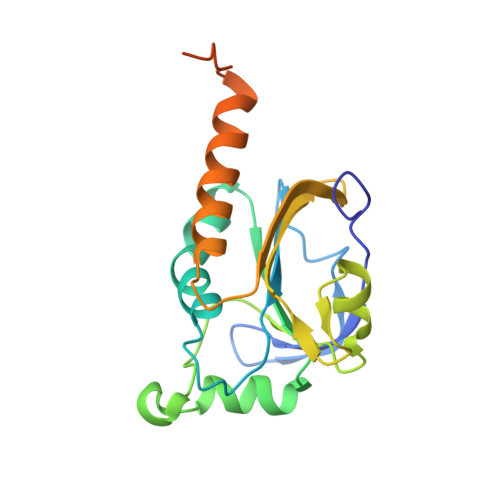
Crystal structure of alkyl hydroperoxide-reductase (AhpC) from Helicobacter pylori.
Papinutto, E., Windle, H.J., Cendron, L., Battistutta, R., Kelleher, D., Zanotti, G.(2005) Biochim Biophys Acta 1753: 240-246
- PubMed: 16213196 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2005.09.001
- Primary Citation Related Structures:
1ZOF - PubMed Abstract:
The AhpC protein from H. pylori, a thioredoxin (Trx)-dependent alkyl hydroperoxide-reductase, is a member of the ubiquitous 2-Cys peroxiredoxins family (2-Cys Prxs), a group of thiol-specific antioxidant enzymes. Prxs exert the protective antioxidant role in cells through their peroxidase activity, whereby hydrogen peroxide, peroxynitrite and a wide range of organic hydroperoxides (ROOH) are reduced and detoxified (ROOH + 2e(-)-->ROH + H2O). In this study AhpC has been cloned and overexpressed in E. coli. After purification to homogeneity, crystals of the recombinant protein were grown. They diffract to 2.95 A resolution using synchrotron radiation. The crystal structure of AhpC has been determined using the molecular replacement method (R = 23.6%, R(free) = 25.9%). The model, similar in the overall to other members of the 2-Cys Prx family crystallized as toroide-shaped complexes, consists of a pentameric arrangement of homodimers [(alpha2)5 decamer]. The model of AhpC from H. pylori presents significant differences with respect to other members of the family: apart from some loop regions, alpha5-helix and the C-terminus is shifted, preventing the C-terminal tail of the second subunit from extending toward this region of the molecule. Oligomerization properties of AhpC have been also characterized by gel filtration chromatography.
- Department of Chemical Sciences, University of Padua, and ICTB, Via Marzolo 1, and Venetian Institute of Molecular Medicine, Via Orus 2, Padua, Italy.
Organizational Affiliation: