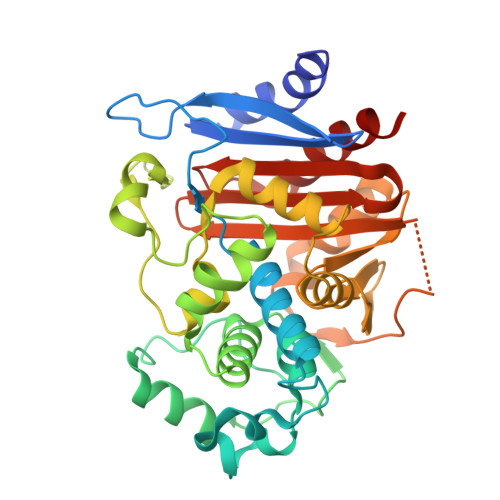
Structural basis for the extended substrate spectrum of CMY-10, a plasmid-encoded class C beta-lactamase.
Kim, J.Y., Jung, H.I., An, Y.J., Lee, J.H., Kim, S.J., Jeong, S.H., Lee, K.J., Suh, P.G., Lee, H.S., Lee, S.H., Cha, S.S.(2006) Mol Microbiol 60: 907-916
- PubMed: 16677302 Search on PubMed
- DOI: https://doi.org/10.1111/j.1365-2958.2006.05146.x
- Primary Citation Related Structures:
1ZKJ - PubMed Abstract:
The emergence and dissemination of extended-spectrum (ES) beta-lactamases induce therapeutic failure and a lack of eradication of clinical isolates even by third-generation beta-lactam antibiotics like ceftazidime. CMY-10 is a plasmid-encoded class C beta-lactamase with a wide spectrum of substrates. Unlike the well-studied class C ES beta-lactamase from Enterobacter cloacae GC1, the Omega-loop does not affect the active site conformation and the catalytic activity of CMY-10. Instead, a three-amino-acid deletion in the R2-loop appears to be responsible for the ES activity of CMY-10. According to the crystal structure solved at 1.55 A resolution, the deletion significantly widens the R2 active site, which accommodates the R2 side-chains of beta-lactam antibiotics. This observation led us to demonstrate the hydrolysing activity of CMY-10 towards imipenem with a long R2 substituent. The forced mutational analyses of P99 beta-lactamase reveal that the introduction of deletion mutations into the R2-loop is able to extend the substrate spectrum of class C non-ES beta-lactamases, which is compatible with the isolation of natural class C ES enzymes harbouring deletion mutations in the R2-loop. Consequently, the opening of the R2 active site by the deletion of some residues in the R2-loop can be considered as an operative molecular strategy of class C beta-lactamases to extend their substrate spectrum.
- School of Biological Sciences, Seoul National University, Seoul 151-742, Republic of Korea.
Organizational Affiliation: