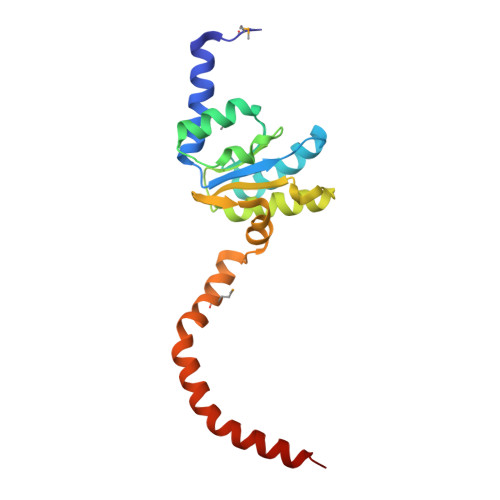
Structural Basis for the Function of the Ribosomal L7/12 Stalk in Factor Binding and GTPase Activation.
Diaconu, M., Kothe, U., Schluenzen, F., Fischer, N., Harms, J.M., Tonevitski, A.G., Stark, H., Rodnina, M.V., Wahl, M.C.(2005) Cell 121: 991-1004
- PubMed: 15989950 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2005.04.015
- Primary Citation Related Structures:
1ZAV, 1ZAW, 1ZAX - PubMed Abstract:
The L7/12 stalk of the large subunit of bacterial ribosomes encompasses protein L10 and multiple copies of L7/12. We present crystal structures of Thermotoga maritima L10 in complex with three L7/12 N-terminal-domain dimers, refine the structure of an archaeal L10E N-terminal domain on the 50S subunit, and identify these elements in cryo-electron-microscopic reconstructions of Escherichia coli ribosomes. The mobile C-terminal helix alpha8 of L10 carries three L7/12 dimers in T. maritima and two in E. coli, in concordance with the different length of helix alpha8 of L10 in these organisms. The stalk is organized into three elements (stalk base, L10 helix alpha8-L7/12 N-terminal-domain complex, and L7/12 C-terminal domains) linked by flexible connections. Highly mobile L7/12 C-terminal domains promote recruitment of translation factors to the ribosome and stimulate GTP hydrolysis by the ribosome bound factors through stabilization of their active GTPase conformation.
- Röntgenkristallographie, Max-Planck-Institut für Biophysikalische Chemie, Am Fassberg 11, D-37077 Göttingen, Germany.
Organizational Affiliation: