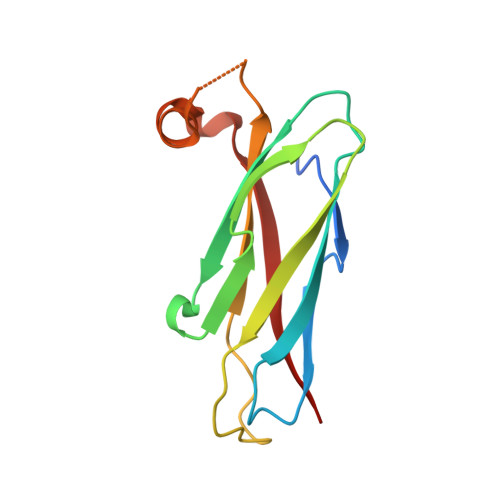
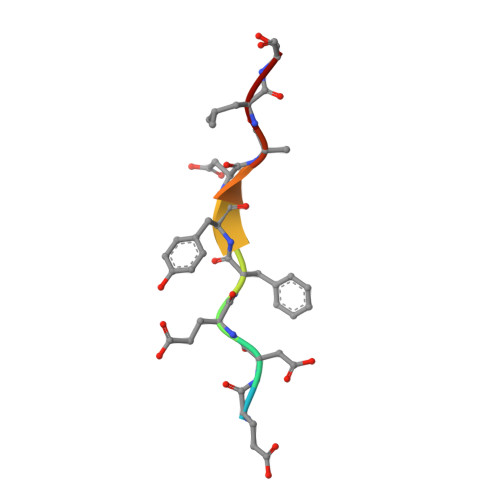
Structural basis of FFAT motif-mediated ER targeting
Kaiser, S.E., Brickner, J.H., Reilein, A.R., Fenn, T.D., Walter, P., Brunger, A.T.(2005) Structure 13: 1035-1045
- PubMed: 16004875 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2005.04.010
- Primary Citation Related Structures:
1Z9L, 1Z9O - PubMed Abstract:
The FFAT motif is a targeting signal responsible for localizing a number of proteins to the cytosolic surface of the endoplasmic reticulum (ER) and to the nuclear membrane. FFAT motifs bind to members of the highly conserved VAP protein family, which are tethered to the cytoplasmic face of the ER by a C-terminal transmembrane domain. We have solved crystal structures of the rat VAP-A MSP homology domain alone and in complex with an FFAT motif. The co-crystal structure was used to design a VAP mutant that disrupts rat and yeast VAP-FFAT interactions in vitro. The FFAT binding-defective mutant also blocked function of the VAP homolog Scs2p in yeast. Finally, overexpression of the FFAT binding-defective VAP in COS7 cells dramatically altered ER morphology. Our data establish the structural basis of FFAT-mediated ER targeting and suggest that FFAT-targeted proteins play an important role in determining ER morphology.
- Department of Molecular and Cellular Physiology, Stanford University, Stanford, CA 94305, USA.
Organizational Affiliation: