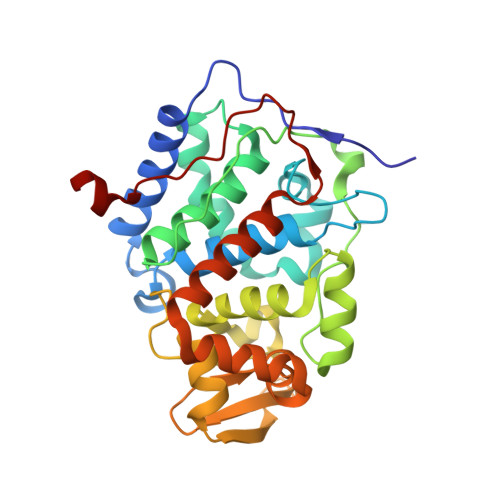
The 1.13-A structure of iron-free cytochrome c peroxidase.
Bhaskar, B., Poulos, T.L.(2005) J Biol Inorg Chem 10: 425-430
- PubMed: 15900441 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-005-0654-4
- Primary Citation Related Structures:
1Z53 - PubMed Abstract:
The iron-free cytochrome c peroxidase (CCP) crystal structure has been determined to 1.13 A and compared with the 1.2-A ferric-CCP structure. Quite unexpectedly, removal of the iron has no effect on porphyrin geometry and distortion, indicating that protein-porphyrin interactions and not iron coordination or formation of the axial His-Fe bond determines porphyrin conformation. However, there are changes in solvent structure in the distal pocket, which lead to changes in the distal His52 acid-base catalyst. The observed ability of His52 to move in response to small changes in solvent structure is very likely important for its role as a catalyst in assisting in the heterolytic fission of the peroxide O-O bond.
- Department of Molecular Biology and Biochemistry and the Center in Chemical and Structural Biology, University of California, Irvine, CA 92697-3900, USA.
Organizational Affiliation: