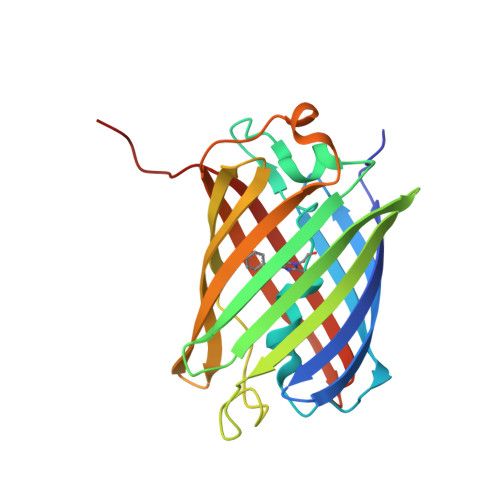
The 2.1A Crystal Structure of the Far-red Fluorescent Protein HcRed: Inherent Conformational Flexibility of the Chromophore
Wilmann, P.G., Petersen, J., Pettikiriarachchi, A., Buckle, A.M., Smith, S.C., Olsen, S., Perugini, M.A., Devenish, R.J., Prescott, M., Rossjohn, J.(2005) J Mol Biology 349: 223-237
- PubMed: 15876379 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.03.020
- Primary Citation Related Structures:
1YZW - PubMed Abstract:
We have determined the crystal structure of HcRed, a far-red fluorescent protein isolated from Heteractis crispa, to 2.1A resolution. HcRed was observed to form a dimer, in contrast to the monomeric form of green fluorescent protein (GFP) or the tetrameric forms of the GFP-like proteins (eqFP611, Rtms5 and DsRed). Unlike the well-defined chromophore conformation observed in GFP and the GFP-like proteins, the HcRed chromophore was observed to be considerably mobile. Within the HcRed structure, the cyclic tripeptide chromophore, Glu(64)-Tyr(65)-Gly(66), was observed to adopt both a cis coplanar and a trans non-coplanar conformation. As a result of these two conformations, the hydroxyphenyl moiety of the chromophore makes distinct interactions within the interior of the beta-can. These data together with a quantum chemical model of the chromophore, suggest the cis coplanar conformation to be consistent with the fluorescent properties of HcRed, and the trans non-coplanar conformation to be consistent with non-fluorescent properties of hcCP, the chromoprotein parent of HcRed. Moreover, within the GFP-like family, it appears that where conformational freedom is permissible then flexibility in the chromophore conformation is possible.
- The Protein Crystallography Unit, Monash Centre for Synchrotron Science, School of Biomedical Sciences, Monash University, Clayton, Vic. 3800, Australia.
Organizational Affiliation: