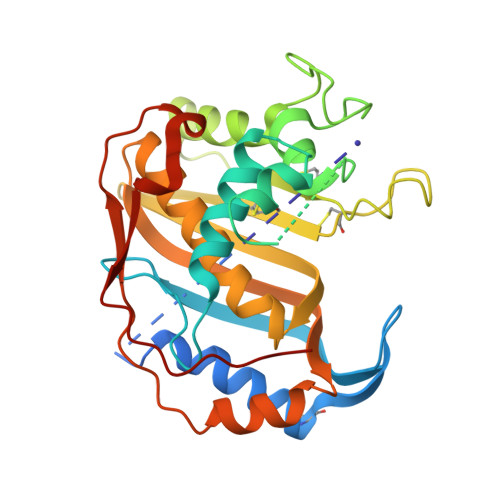
Structure of human thymidylate synthase under low-salt conditions.
Lovelace, L.L., Minor, W., Lebioda, L.(2005) Acta Crystallogr D Biol Crystallogr 61: 622-627
- PubMed: 15858273 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444905005895
- Primary Citation Related Structures:
1YPV - PubMed Abstract:
Human thymidylate synthase, a target in cancer chemotherapy, was crystallized from PEG 3350 with 30 mM ammonium sulfate (AS) in the crystallization medium. The crystals are isomorphous with the high-salt crystals ( approximately 2.0 M AS) and the structure has been solved and refined (R = 22.6%, R(free) = 24.3%) at 1.8 A resolution. The high- and low-AS-concentration structures are quite similar, with loop 181-197 is in the inactive conformation. Also, residues 95-106 and 129-135 (eukaryotic inserts region) show high mobility as assessed by poor electron density and high values of crystallographic temperature factors (residues 1-25 and 108-129 are disordered in both structures). The high mobility of this region may reflect the situation at physiological ionic strength. Of the four sulfate ions observed bound at 2.0 M AS, only two are present at 30 mM AS. The inactive conformation appears to be stabilized by the side chain of Val3 or a leucine residue from the disordered regions. The low-salt conditions of these crystals should be much more suitable for the study of thymidylate synthase inhibitors, especially those that utilize sulfate-binding sites to stabilize the inactive conformation of loop 181-197.
- Department of Chemistry and Biochemistry, University of South Carolina, Columbia, SC 29208, USA.
Organizational Affiliation: