Crystal structure of a non-hemorrhagic fibrin(ogen)olytic metalloproteinase complexed with a novel natural tri-peptide inhibitor from venom of Agkistrodon acutus
Lou, Z., Hou, J., Liang, X., Chen, J., Qiu, P., Liu, Y., Li, M., Rao, Z., Yan, G.(2005) J Struct Biol 152: 195-203
- PubMed: 16330227 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2005.09.006
- Primary Citation Related Structures:
1YP1 - PubMed Abstract:
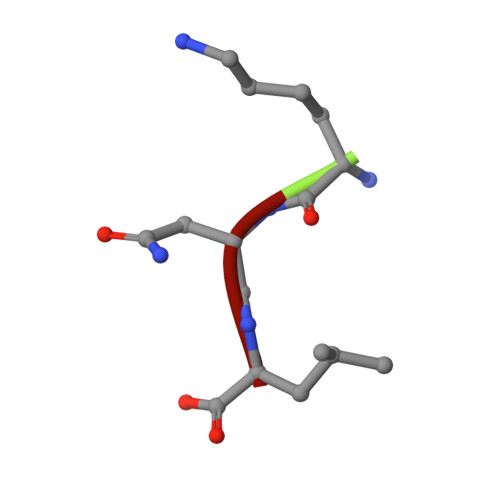
Thrombotic occlusive diseases pose a great threat to human health. Thrombolytic agents are in widespread use for the dissolution of arterial and venous pathologic thrombi in these kinds of diseases. Snake venom metalloproteinases (SVMPs) can act directly on fibrin/fibrinogen and are therefore potential candidates for therapeutic use against thrombotic occlusive diseases. In this study, we have determined the crystal structure of FII, a novel non-hemorrhagic SVMP isolated from Anhui Agkistrodon acutus snake venom by molecular replacement. The structure reveals that FII is a member of the P-I class SVMPs. The Zn2+ ion essential for hydrolytic activity is found in the active site and is tetrahedrally co-ordinated by three histidine residues and water molecule. Unambiguous electron density for a tri-peptide with sequence KNL is also found located near the active site. Biochemical evidences show that the tri-peptide KNL can inhibit the enzymatic activity of FII.
- Laboratory of Structural Biology, Tsinghua University, Beijing 100084, People's Republic of China.
Organizational Affiliation: