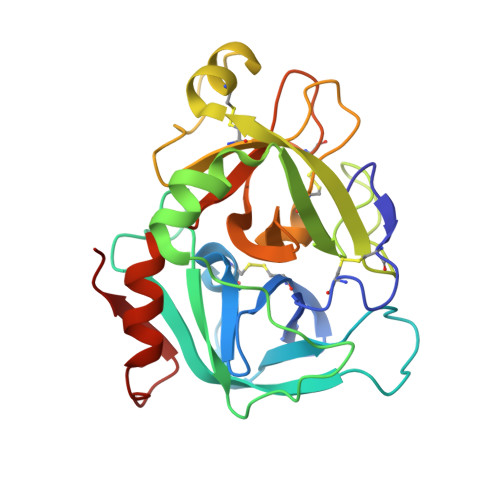
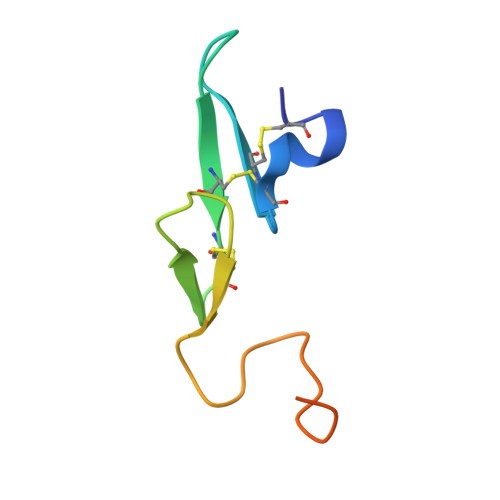
A selective, slow binding inhibitor of factor VIIa binds to a nonstandard active site conformation and attenuates thrombus formation in vivo.
Olivero, A.G., Eigenbrot, C., Goldsmith, R., Robarge, K., Artis, D.R., Flygare, J., Rawson, T., Sutherlin, D.P., Kadkhodayan, S., Beresini, M., Elliott, L.O., DeGuzman, G.G., Banner, D.W., Ultsch, M., Marzec, U., Hanson, S.R., Refino, C., Bunting, S., Kirchhofer, D.(2005) J Biological Chem 280: 9160-9169
- PubMed: 15632123 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M409068200
- Primary Citation Related Structures:
1YGC - PubMed Abstract:
The serine protease factor VIIa (FVIIa) in complex with its cellular cofactor tissue factor (TF) initiates the blood coagulation reactions. TF.FVIIa is also implicated in thrombosis-related disorders and constitutes an appealing therapeutic target for treatment of cardiovascular diseases. To this end, we generated the FVIIa active site inhibitor G17905, which displayed great potency toward TF.FVIIa (Ki = 0.35 +/- 0.11 nM). G17905 did not appreciably inhibit 12 of the 14 examined trypsin-like serine proteases, consistent with its TF.FVIIa-specific activity in clotting assays. The crystal structure of the FVIIa.G17905 complex provides insight into the molecular basis of the high selectivity. It shows that, compared with other serine proteases, FVIIa is uniquely equipped to accommodate conformational disturbances in the Gln217-Gly219 region caused by the ortho-hydroxy group of the inhibitor's aminobenzamidine moiety located in the S1 recognition pocket. Moreover, the structure revealed a novel, nonstandard conformation of FVIIa active site in the region of the oxyanion hole, a "flipped" Lys192-Gly193 peptide bond. Macromolecular substrate activation assays demonstrated that G17905 is a noncompetitive, slow-binding inhibitor. Nevertheless, G17905 effectively inhibited thrombus formation in a baboon arterio-venous shunt model, reducing platelet and fibrin deposition by approximately 70% at 0.4 mg/kg + 0.1 mg/kg/min infusion. Therefore, the in vitro potency of G17905, characterized by slow binding kinetics, correlated with efficacious antithrombotic activity in vivo.
- Department of Medicinal Chemistry, Genentech, Inc., South San Francisco, California 94080, USA.
Organizational Affiliation: