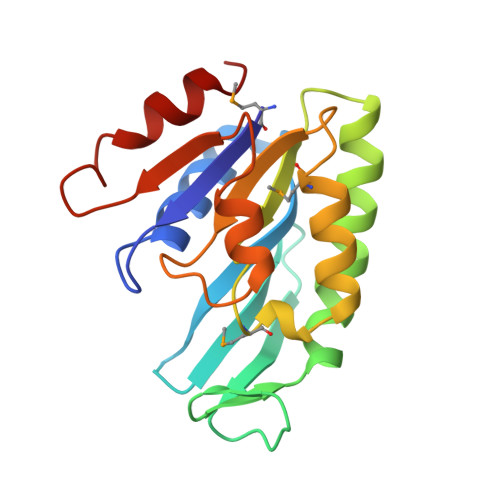
Crystal structure of THEP1 from the hyperthermophile Aquifex aeolicus: a variation of the RecA fold
Rossbach, M., Daumke, O., Klinger, C., Wittinghofer, A., Kaufmann, M.(2005) BMC Struct Biol 5: 7-7
- PubMed: 15777481 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-5-7
- Primary Citation Related Structures:
1YE8 - PubMed Abstract:
aaTHEP1, the gene product of aq_1292 from Aquifex aeolicus, shows sequence homology to proteins from most thermophiles, hyperthermophiles, and higher organisms such as man, mouse, and fly. In contrast, there are almost no homologous proteins in mesophilic unicellular microorganisms. aaTHEP1 is a thermophilic enzyme exhibiting both ATPase and GTPase activity in vitro. Although annotated as a nucleotide kinase, such an activity could not be confirmed for aaTHEP1 experimentally and the in vivo function of aaTHEP1 is still unknown. Here we report the crystal structure of selenomethionine substituted nucleotide-free aaTHEP1 at 1.4 A resolution using a multiple anomalous dispersion phasing protocol. The protein is composed of a single domain that belongs to the family of 3-layer (alpha/beta/alpha)-structures consisting of nine central strands flanked by six helices. The closest structural homologue as determined by DALI is the RecA family. In contrast to the latter proteins, aaTHEP1 possesses an extension of the beta-sheet consisting of four additional beta-strands. We conclude that the structure of aaTHEP1 represents a variation of the RecA fold. Although the catalytic function of aaTHEP1 remains unclear, structural details indicate that it does not belong to the group of GTPases, kinases or adenosyltransferases. A mainly positive electrostatic surface indicates that aaTHEP1 might be a DNA/RNA modifying enzyme. The resolved structure of aaTHEP1 can serve as paradigm for the complete THEP1 family.
- Institute of Neurobiochemistry, The Protein Chemistry Group, Witten/Herdecke University, Stockumer Strasse 10, 58448 Witten, Germany. miro@uni-wh.de
Organizational Affiliation: