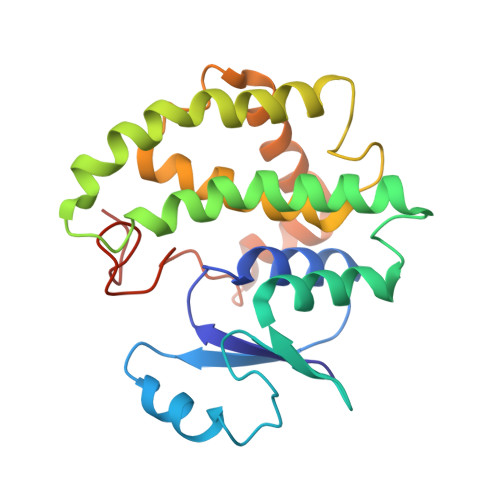
X-ray structure of glutathione S-transferase from Schistosoma japonicum in a new crystal form reveals flexibility of the substrate-binding site
Rufer, A.C., Thiebach, L., Baer, K., Klein, H.W., Hennig, M.(2005) Acta Crystallogr Sect F Struct Biol Cryst Commun 61: 263-265
- PubMed: 16511012 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309105004823
- Primary Citation Related Structures:
1Y6E - PubMed Abstract:
The crystal structure of the 26 kDa glutathione S-transferase from Schistosoma japonicum (SjGST) was determined at 3 A resolution in the new space group P2(1)2(1)2(1). The structure of orthorhombic SjGST reveals unique features of the ligand-binding site and dimer interface when compared with previously reported structures. SjGST is recognized as the major detoxification enzyme of S. japonicum, a pathogenic helminth causing schistosomiasis. As resistance against the established inhibitor of SjGST, praziquantel, has been reported these results might prove to be valuable for the development of novel drugs.
- F. Hoffmann-La Roche AG, Pharma Research Discovery Chemistry, 4070 Basel, Switzerland. arne.rufer@roche.com
Organizational Affiliation: