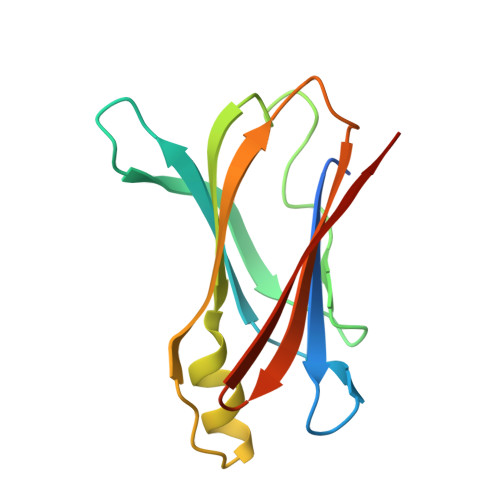
Human transthyretin in complex with iododiflunisal: structural features associated with a potent amyloid inhibitor.
Gales, L., Macedo-Ribeiro, S., Arsequell, G., Valencia, G., Saraiva, M.J., Damas, A.M.(2005) Biochem J 388: 615-621
- PubMed: 15689188 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20042035
- Primary Citation Related Structures:
1Y1D - PubMed Abstract:
Ex vivo and in vitro studies have revealed the remarkable amyloid inhibitory potency and specificity of iododiflunisal in relation to transthyretin [Almeida, Macedo, Cardoso, Alves, Valencia, Arsequell, Planas and Saraiva (2004) Biochem. J. 381, 351-356], a protein implicated in familial amyloidotic polyneuropathy. In the present paper, the crystal structure of transthyretin complexed with this diflunisal derivative is reported, which enables a detailed analysis of the protein-ligand interactions. Iododiflunisal binds very deep in the hormone-binding channel. The iodine substituent is tightly anchored into a pocket of the binding site and the fluorine atoms provide extra hydrophobic contacts with the protein. The carboxylate substituent is involved in an electrostatic interaction with the N(zeta) of a lysine residue. Moreover, ligand-induced conformational alterations in the side chain of some residues result in the formation of new intersubunit hydrogen bonds. All these new interactions, induced by iododiflunisal, increase the stability of the tetramer impairing the formation of amyloid fibrils. The crystal structure of this complex opens perspectives for the design of more specific and effective drugs for familial amyloidotic polyneuropathy patients.
- Instituto de Ciências Biomédicas Abel Salazar and Instituto de Biologia Molecular e Celular, Rua do Campo Alegre 823, 4150 Porto, Portugal.
Organizational Affiliation: