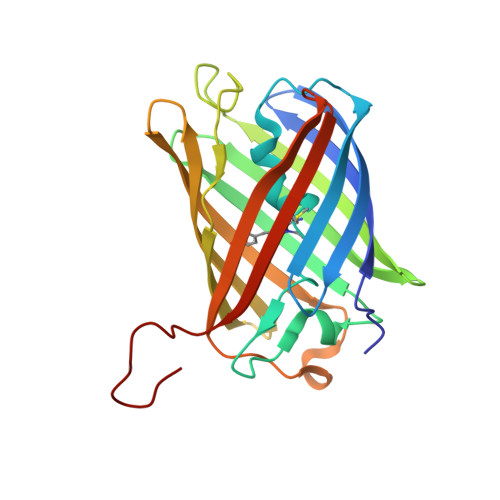
Variations on the GFP chromophore: A polypeptide fragmentation within the chromophore revealed in the 2.1-A crystal structure of a nonfluorescent chromoprotein from Anemonia sulcata
Wilmann, P.G., Petersen, J., Devenish, R.J., Prescott, M., Rossjohn, J.(2005) J Biological Chem 280: 2401-2404
- PubMed: 15542608 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.C400484200
- Primary Citation Related Structures:
1XQM - PubMed Abstract:
We have determined to 2.1 A resolution the crystal structure of a dark state, kindling fluorescent protein isolated from the sea anemone, Anemonia sulcata. The chromophore sequence Met(63)-Tyr(64)-Gly(65) of the A. sulcata chromoprotein was previously proposed to comprise a 6-membered pyrazine-type heterocycle (Martynov, V. I., Savitsky, A. P., Martynova, N. Y., Savitsky, P. A., Lukyanov, K. A., and Lukyanov, S. A. (2001) J. Biol. Chem. 276, 21012-21016). However, our crystallographic data revealed the chromophore to comprise a 5-membered p-hydroxybenzylideneimidazolinone moiety that adopts a non-coplanar trans conformation within the interior of the GFP beta-can fold. Unexpectedly, fragmentation of the polypeptide was found to occur within the chromophore moiety, at the bond between Cys(62C) and Met(63N1.) Our structural data reveal that fragmentation of the chromophore represents an intrinsic, autocatalytic step toward the formation of the mature chromophore within the specific GFP-like proteins.
- Protein Crystallography Unit, Monash Centre for Synchrotron Science.
Organizational Affiliation: