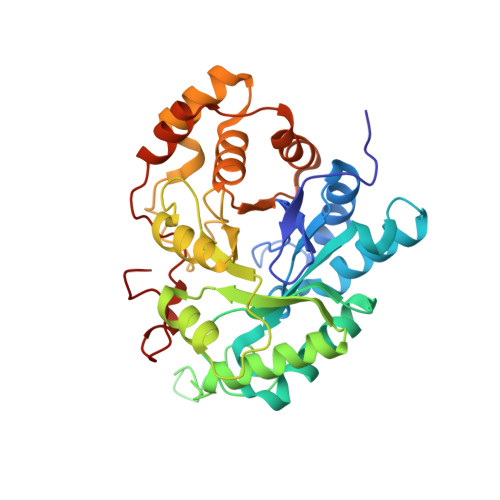
Comparison of crystal structures of human type 3 3alpha-hydroxysteroid dehydrogenase reveals an "induced-fit" mechanism and a conserved basic motif involved in the binding of androgen
Couture, J.-F., Roy, A.-M., Cantin, L., Legrand, P., Luu-The, V., Labrie, F., Breton, R.(2005) Protein Sci 14: 1485-1497
- PubMed: 15929998 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.051353205
- Primary Citation Related Structures:
1XJB - PubMed Abstract:
The aldo-keto reductase (AKR) human type 3 3alpha-hydroxysteroid dehydrogenase (h3alpha-HSD3, AKR1C2) plays a crucial role in the regulation of the intracellular concentrations of testosterone and 5alpha-dihydrotestosterone (5alpha-DHT), two steroids directly linked to the etiology and the progression of many prostate diseases and cancer. This enzyme also binds many structurally different molecules such as 4-hydroxynonenal, polycyclic aromatic hydrocarbons, and indanone. To understand the mechanism underlying the plasticity of its substrate-binding site, we solved the binary complex structure of h3alpha-HSD3-NADP(H) at 1.9 A resolution. During the refinement process, we found acetate and citrate molecules deeply engulfed in the steroid-binding cavity. Superimposition of this structure with the h3alpha-HSD3-NADP(H)-testosterone/acetate ternary complex structure reveals that one of the mobile loops forming the binding cavity operates a slight contraction movement against the citrate molecule while the side chains of many residues undergo numerous conformational changes, probably to create an optimal binding site for the citrate. These structural changes, which altogether cause a reduction of the substrate-binding cavity volume (from 776 A(3) in the presence of testosterone/acetate to 704 A(3) in the acetate/citrate complex), are reminiscent of the "induced-fit" mechanism previously proposed for the aldose reductase, another member of the AKR superfamily. We also found that the replacement of residues Arg(301) and Arg(304), localized near the steroid-binding cavity, significantly affects the 3alpha-HSD activity of this enzyme toward 5alpha-DHT and completely abolishes its 17beta-HSD activity on 4-dione. All these results have thus been used to reevaluate the binding mode of this enzyme for androgens.
- Oncology and Molecular Endocrinology Research Center, Laval University Medical Center and Laval University, Québec, Canada.
Organizational Affiliation: