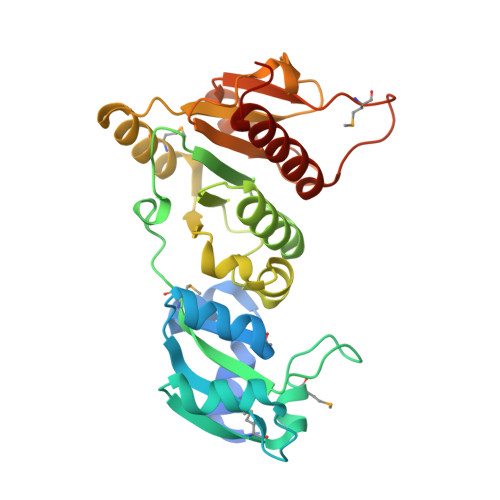
Structure and function of the antibiotic resistance-mediating methyltransferase AviRb from Streptomyces viridochromogenes
Mosbacher, T.G., Bechthold, A., Schulz, G.E.(2005) J Mol Biology 345: 535-545
- PubMed: 15581897 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.10.051
- Primary Citation Related Structures:
1X7O, 1X7P - PubMed Abstract:
The emergence of antibiotic-resistant bacterial strains is a widespread problem in medical practice and drug design, and each case requires the elucidation of the underlying mechanism. AviRb from Streptomyces viridochromogenes methylates the 2'-O atom of U2479 of the 23S ribosomal RNA in Gram-positive bacteria and thus mediates resistance to the oligosaccharide (orthosomycin) antibiotic avilamycin. The structure of AviRb with and without bound cofactor S-adenosyl-L-methionine (AdoMet) was determined, showing that it is a homodimer belonging to the SpoU family within the SPOUT class of methyltransferases. The relationships within this class were analyzed in detail and, in addition, a novel fourth SpoU sequence fingerprint is proposed. Each subunit of AviRb consists of two domains. The N-terminal domain, being related to the ribosomal proteins L30 and L7Ae, is likely to bind RNA. The C-terminal domain is related to all SPOUT methyltransferases, and is responsible for AdoMet-binding, catalysis and dimerization. The cofactor binds at the characteristic knot of the polypeptide in an unusually bent conformation. The transferred methyl group points to a broad cleft formed with the L30-type domain of the other subunit. Measurements of mutant activity revealed four important residues responsible for catalysis and allowed the modeling of a complex between AviRb and the RNA target. The model includes a specificity pocket for uracil but does not contain a base for deprotonating the 2'-O atom of U2479 on methylation.
- Institut für Organische Chemie und Biochemie, Albert-Ludwigs-Universität, Albertstr. 21, D-79104 Freiburg im Breisgau, Germany.
Organizational Affiliation: