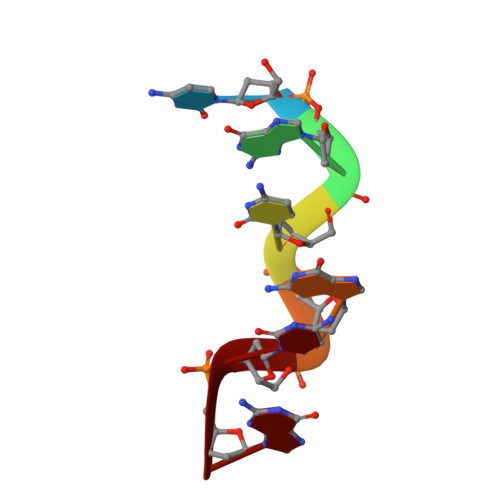
The hydration structure of a Z-DNA hexameric duplex determined by a neutron diffraction technique.
Chatake, T., Tanaka, I., Umino, H., Arai, S., Niimura, N.(2005) Acta Crystallogr D Biol Crystallogr 61: 1088-1098
- PubMed: 16041074 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444905015581
- Primary Citation Related Structures:
1V9G, 1WOE - PubMed Abstract:
In order to reveal the hydration structure of Z-DNA, a neutron diffraction study has been carried out at 1.8 A resolution on a Z-DNA hexamer d(CGCGCG). Neutron diffraction data were collected with the BIX-3 single-crystal diffractometer at the JRR-3 reactor in the Japan Atomic Energy Research Institute (JAERI) using a large crystal (1.6 mm3) obtained from D2O solution. It has been found that almost all the guanine bases have participated in H/D exchange at the C8-H8 group, consistent with the acidic nature of this bond. 44 water molecules were found in the nuclear density maps, of which 29 showed the entire contour of all three atoms (D-O-D). The remaining 15 water molecules had a simple spherical shape, indicating that they were rotationally disordered. An interesting relationship was found between the orientational disorder of the water molecules and their locations. Almost all water molecules in the minor groove were well ordered in the crystal, while 40% of the water molecules in the major groove were rotationally disordered. The hydrogen-bonding networks in the hydration shells have two structural aspects: flexibility and regularity.
- Faculty of Pharmaceutical Sciences, Chiba Institute of Science, Choshi, Chiba 288-0025, Japan.
Organizational Affiliation: