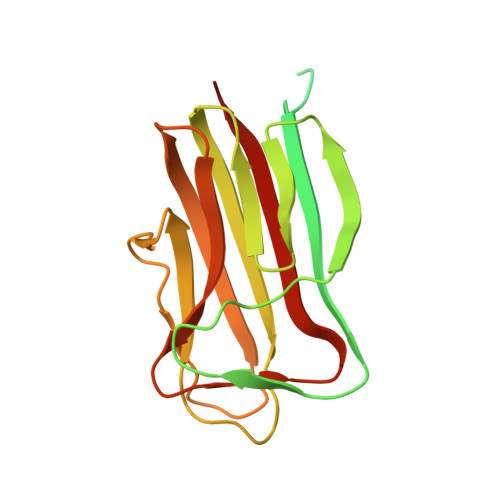
The Crystal Structure of the Bacillus Anthracis Spore Surface Protein Bcla Shows Remarkable Similarity to Mammalian Proteins.
Rety, S., Salamitou, S., Garcia-Verdugo, I., Hulmes, D.J., Lehegarat, F., Chaby, R., Lewit-Bentley, A.(2005) J Biological Chem 280: 43073
- PubMed: 16249180 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M510087200
- Primary Citation Related Structures:
1WCK - PubMed Abstract:
The lethal disease anthrax is propagated by spores of Bacillus anthracis, which can penetrate into the mammalian host by inhalation, causing a rapid progression of the disease and a mostly fatal outcome. We have solved the three-dimensional structure of the major surface protein BclA on B. anthracis spores. Surprisingly, the structure resembles C1q, the first component of complement, despite there being no sequence homology. Although most assays for C1q-like activity, including binding to C1q receptors, suggest that BclA does not mimic C1q, we show that BclA, as well as C1q, interacts with components of the lung alveolar surfactant layer. Thus, to better recognize and invade its hosts, this pathogenic soil bacterium may have evolved a surface protein whose structure is strikingly close to a mammalian protein.
- Laboratoire de Biotechnologies et Pharmacologie Génétique Appliquées, CNRS, Unité Mixte de Recherche 8113, Ecole Normale Supérieure de Cachan, 61 Avenue du Président Wilson, 94235 Cachan, France.
Organizational Affiliation: