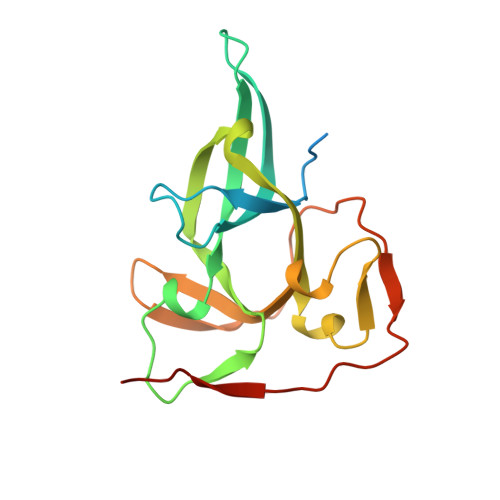
Crystal Structure of the Proximal Bah Domain of the Polybromo Protein
Oliver, A.W., Jones, S.A., Roe, S.M., Matthews, S., Goodwin, G.H., Pearl, L.H.(2005) Biochem J 389: 657
- PubMed: 15839835 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20050310
- Primary Citation Related Structures:
1W4S - PubMed Abstract:
The BAH domain (bromo-associated homology domain) was first identified from a repeated motif found in the nuclear protein polybromo--a large (187 kDa) modular protein comprising six bromodomains, two BAH domains and an HMG box. To date, the BAH domain has no ascribed function, although it is found in a wide range of proteins that contain additional domains involved in either transcriptional regulation (e.g. SET, PHD and bromodomain) and/or DNA binding (HMG box and AT hook). The molecular function of polybromo itself also remains unclear, but it has been identified as a key component of an SWI/SNF (switching/sucrose non-fermenting)-related, ATP-dependent chromatin-remodelling complex PBAF (polybromo, BRG1-associated factors; also known as SWI/SNF-B or SWI/SNFbeta). We present in this paper the crystal structure of the proximal BAH domain from chicken polybromo (BAH1), at a resolution of 1.6 A (1 A=0.1 nm). Structure-based sequence analysis reveals several features that may be involved in mediating protein-protein interactions.
- CR-UK DNA Repair Enzymes Group, Section of Structural Biology, The Institute of Cancer Research, 237 Fulham Road, Chelsea, London SW3 6JB, UK. antony.oliver@icr.ac.uk
Organizational Affiliation: