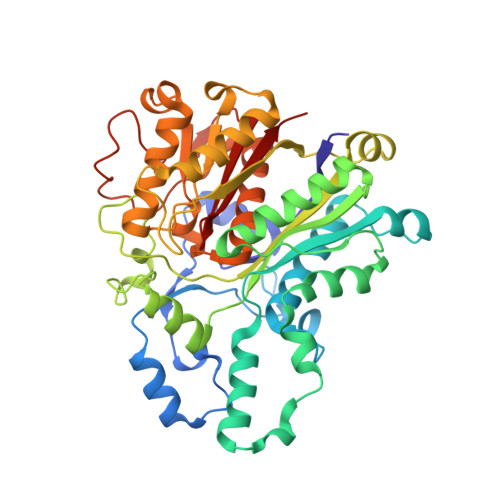
Structure of the mitochondrial beta-ketoacyl-[acyl carrier protein] synthase from Arabidopsis and its role in fatty acid synthesis.
Olsen, J.G., Rasmussen, A.V., von Wettstein-Knowles, P., Henriksen, A.(2004) FEBS Lett 577: 170-174
- PubMed: 15527780 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2004.10.007
- Primary Citation Related Structures:
1W0I - PubMed Abstract:
Mitochondrial fatty acid synthesis is catalyzed by a dissociated fatty acid synthase similar to those of plant plastids and bacteria. The crystal structure of a mitochondrial beta-ketoacyl-[acyl carrier protein] synthase (mtKAS), namely that from Arabidopsis thaliana, has been determined for the first time. This enzyme accomplishes the vital condensation steps in constructing fatty acid carbon skeletons. The product profile of mtKAS is unusual in that C8 and C(14-16) fatty acyl chains predominate. An enzyme architecture that likely is the basis for the observed bimodal profile of mtKAS products can be derived from the shape of the acyl binding pocket.
- Carlsberg Laboratory, Gamle Carlsberg Vej 10, DK-2500 Valby, Denmark.
Organizational Affiliation: