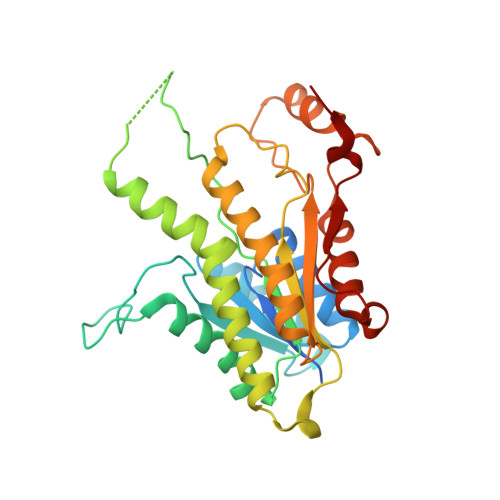
Inhibition of Leishmania Major Pteridine Reductase by 2,4,6-Triaminoquinazoline: Structure of the Nadph Ternary Complex
Mcluskey, K., Gibellini, F., Carvalho, P., Avery, M., Hunter, W.(2004) Acta Crystallogr D Biol Crystallogr 60: 1780
- PubMed: 15388924 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904018955
- Primary Citation Related Structures:
1W0C - PubMed Abstract:
The structure of Leishmania major pteridine reductase (PTR1) in complex with NADPH and the inhibitor 2,4,6-triaminoquinazoline (TAQ) has been solved in a new crystal form by molecular replacement and refined to 2.6 A resolution. The inhibitor mimics a fragment, the pterin head group, of the archetypal antifolate drug methotrexate (MTX) and exploits similar chemical features to bind in the PTR1 active site. Despite being a much smaller molecule, TAQ displays a similar inhibition constant to that of MTX. PTR1 is a target for the development of improved therapies for infections caused by trypanosomatid parasites and this analysis provides information to assist the structure-based development of novel enzyme inhibitors.
- Division of Biological Chemistry and Molecular Microbiology, School of Life Sciences, University of Dundee, Dundee DD1 5EH, Scotland.
Organizational Affiliation: