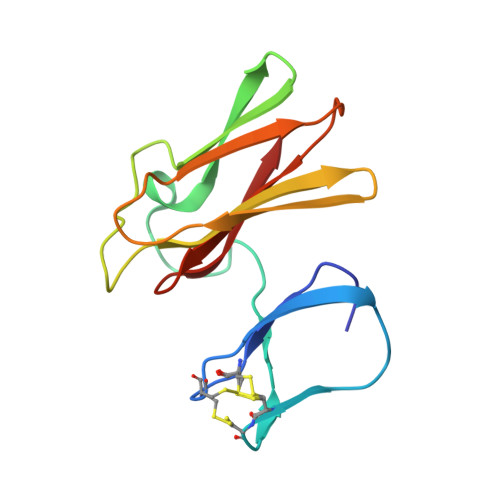
Structure of Superoxide Reductase Bound to Ferrocyanide and Active Site Expansion Upon X-Ray Induced Photoreduction
Adam, V., Royant, A., Niviere, V., Molina-Heredia, F.P., Bourgeois, D.(2004) Structure 12: 1729
- PubMed: 15341736 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.07.013
- Primary Citation Related Structures:
1VZG, 1VZH, 1VZI - PubMed Abstract:
Some sulfate-reducing and microaerophilic bacteria rely on the enzyme superoxide reductase (SOR) to eliminate the toxic superoxide anion radical (O2*-). SOR catalyses the one-electron reduction of O2*- to hydrogen peroxide at a nonheme ferrous iron center. The structures of Desulfoarculus baarsii SOR (mutant E47A) alone and in complex with ferrocyanide were solved to 1.15 and 1.7 A resolution, respectively. The latter structure, the first ever reported of a complex between ferrocyanide and a protein, reveals that this organo-metallic compound entirely plugs the SOR active site, coordinating the active iron through a bent cyano bridge. The subtle structural differences between the mixed-valence and the fully reduced SOR-ferrocyanide adducts were investigated by taking advantage of the photoelectrons induced by X-rays. The results reveal that photo-reduction from Fe(III) to Fe(II) of the iron center, a very rapid process under a powerful synchrotron beam, induces an expansion of the SOR active site.
- LCCP, UMR 5075, IBS-CEA/CNRS/Université J. Fourier, 41 Avenue Jules Horowitz, 38027 Grenoble, Cedex 1, France.
Organizational Affiliation: