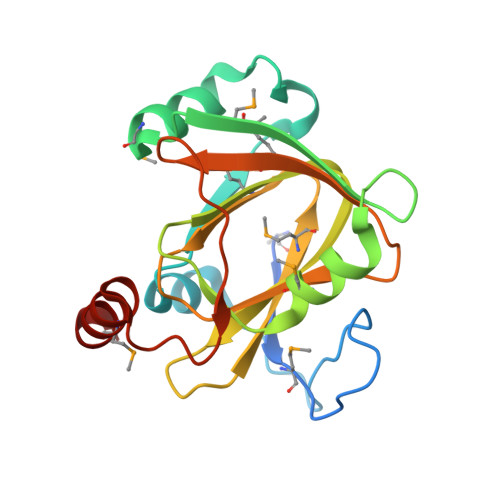
Crystal structure of acireductone dioxygenase (ARD) from Mus musculus at 2.06 angstrom resolution.
Xu, Q., Schwarzenbacher, R., Krishna, S.S., McMullan, D., Agarwalla, S., Quijano, K., Abdubek, P., Ambing, E., Axelrod, H., Biorac, T., Canaves, J.M., Chiu, H.J., Elsliger, M.A., Grittini, C., Grzechnik, S.K., DiDonato, M., Hale, J., Hampton, E., Han, G.W., Haugen, J., Hornsby, M., Jaroszewski, L., Klock, H.E., Knuth, M.W., Koesema, E., Kreusch, A., Kuhn, P., Miller, M.D., Moy, K., Nigoghossian, E., Paulsen, J., Reyes, R., Rife, C., Spraggon, G., Stevens, R.C., van den Bedem, H., Velasquez, J., White, A., Wolf, G., Hodgson, K.O., Wooley, J., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2006) Proteins 64: 808-813
- PubMed: 16783794 Search on PubMed
- DOI: https://doi.org/10.1002/prot.20947
- Primary Citation Related Structures:
1VR3 - The Joint Center for Structural Genomics, Stanford University, Menlo Park, California, USA.
Organizational Affiliation: