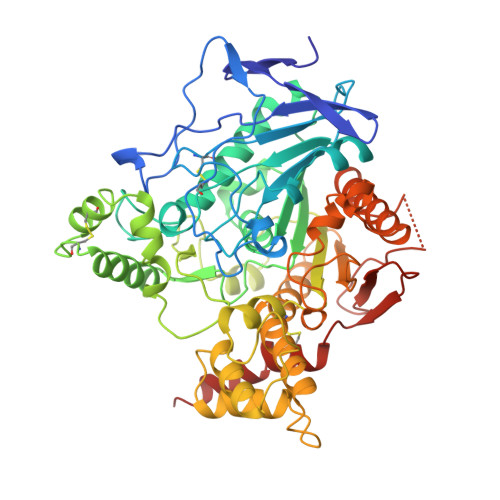
Structure of acetylcholinesterase complexed with the nootropic alkaloid, (-)-huperzine A.
Raves, M.L., Harel, M., Pang, Y.P., Silman, I., Kozikowski, A.P., Sussman, J.L.(1997) Nat Struct Biol 4: 57-63
- PubMed: 8989325 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0197-57
- Primary Citation Related Structures:
1VOT, 2ACE - PubMed Abstract:
(-)-Huperzine A (HupA) is found in an extract from a club moss that has been used for centuries in Chinese folk medicine. Its action has been attributed to its ability to strongly inhibit acetylcholinesterase (AChE). The crystal structure of the complex of AChE with optically pure HupA at 2.5 A resolution shows an unexpected orientation for the inhibitor with surprisingly few strong direct interactions with protein residues to explain its high affinity. This structure is compared to the native structure of AChE devoid of any inhibitor as determined to the same resolution. An analysis of the affinities of structural analogues of HupA, correlated with their interactions with the protein, shows the importance of individual hydrophobic interactions between HupA and aromatic residues in the active-site gorge of AChE.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot, Israel.
Organizational Affiliation: