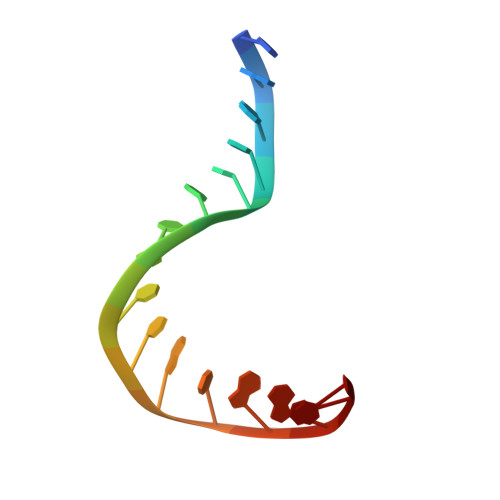
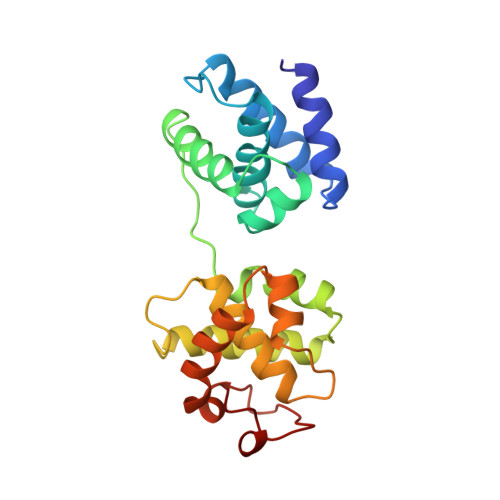
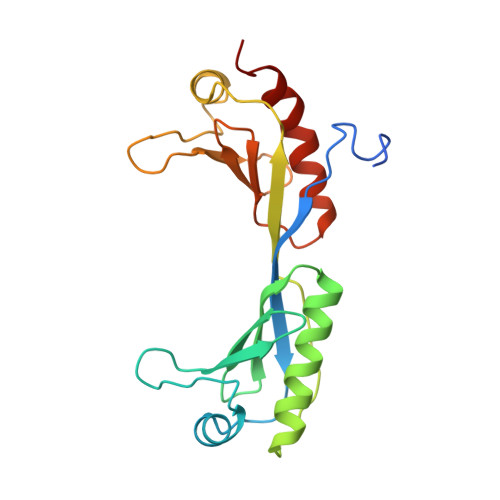
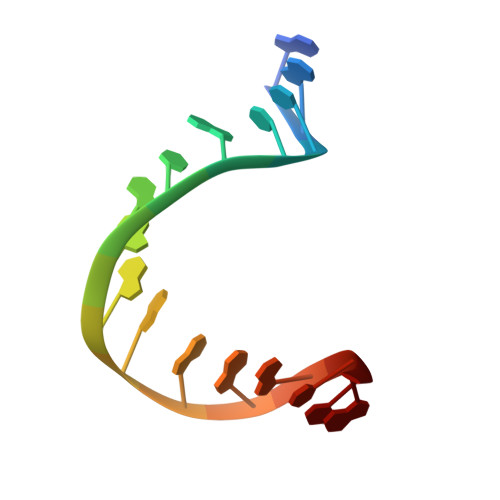
Crystal structure of a TFIIB-TBP-TATA-element ternary complex.
Nikolov, D.B., Chen, H., Halay, E.D., Usheva, A.A., Hisatake, K., Lee, D.K., Roeder, R.G., Burley, S.K.(1995) Nature 377: 119-128
- PubMed: 7675079 Search on PubMed
- DOI: https://doi.org/10.1038/377119a0
- Primary Citation Related Structures:
1VOL - PubMed Abstract:
The crystal structure of the transcription factor IIB (TFIIB)/TATA box-binding protein (TBP)/TATA-element ternary complex is described at 2.7 A resolution. Core TFIIB resembles cyclin A, and recognizes the preformed TBP-DNA complex through protein-protein and protein-DNA interactions. The amino-terminal domain of core TFIIB forms the downstream surface of the ternary complex, where it could fix the transcription start site. The remaining surfaces of TBP and the TFIIB can interact with TBP-associated factors, other class II initiation factors, and transcriptional activators and coactivators.
- Laboratories of Molecular Biophysics, Rockefeller University, New York, New York 10021, USA.
Organizational Affiliation: