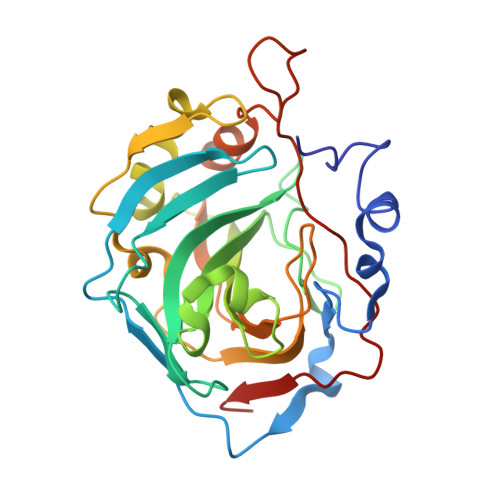
Structure of bovine carbonic anhydrase II at 1.95 A resolution.
Saito, R., Sato, T., Ikai, A., Tanaka, N.(2004) Acta Crystallogr D Biol Crystallogr 60: 792-795
- PubMed: 15039588 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904003166
- Primary Citation Related Structures:
1V9E - PubMed Abstract:
Carbonic anhydrase (CA) is a zinc-containing enzyme that catalyzes the reversible hydration of CO2 to HCO3-. In eukaryotes, the enzyme plays a role in various physiological functions, including interconversion between CO2 and HCO3- in intermediary metabolism, facilitated diffusion of CO2, pH homeostasis and ion transport. The structure of bovine carbonic anhydrase II (BCA II) has been determined by molecular replacement and refined to 1.95 A resolution by simulated-annealing and individual B-factor refinement. The final R factor for the BCA II structure was 19.4%. BCA II has a C-terminal knot structure similar to that observed in human CA II. It contains one zinc ion in the active site coordinated to three histidines and one putative water molecule in a tetrahedral geometry. The structure of BCA II reveals a probable alternative proton-wire pathway that differs from that of HCA II.
- Department of Life Science, Graduate School of Bioscience and Biotechnology, Tokyo Institute of Technology, Japan.
Organizational Affiliation: