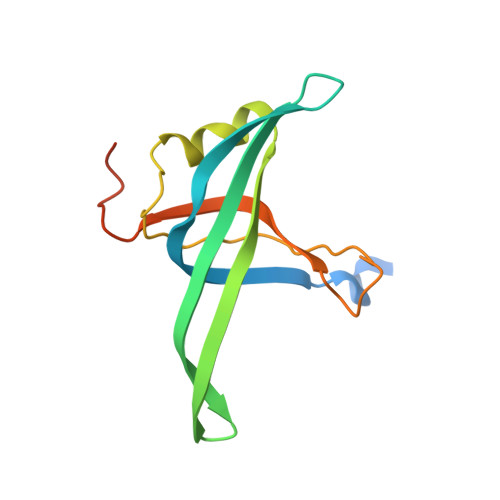
Crystal Structure of Prib- a Primosomal DNA Replication Protein of Escherichia Coli
Liu, J.-H., Chang, T.-W., Huang, C.-Y., Chen, S.-U., Wu, H.-N., Chang, M.-C., Hsiao, C.-D.(2004) J Biological Chem 279: 50465
- PubMed: 15383524 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M406773200
- Primary Citation Related Structures:
1V1Q - PubMed Abstract:
PriB is one of the Escherichia coli varphiX-type primosome proteins that are required for assembly of the primosome, a mobile multi-enzyme complex responsible for the initiation of DNA replication. Here we report the crystal structure of the E. coli PriB at 2.1 A resolution by multi-wavelength anomalous diffraction using a mercury derivative. The polypeptide chain of PriB is structurally similar to that of single-stranded DNA-binding protein (SSB). However, the biological unit of PriB is a dimer, not a homotetramer like SSB. Electrophoretic mobility shift assays demonstrated that PriB binds single-stranded DNA and single-stranded RNA with comparable affinity. We also show that PriB binds single-stranded DNA with certain base preferences. Based on the PriB structural information and biochemical studies, we propose that the potential tetramer formation surface and several other regions of PriB may participate in protein-protein interaction during DNA replication. These findings may illuminate the role of PriB in varphiX-type primosome assembly.
- Graduate Institute of Life Sciences, National Defense Medical Center, Taipei, Taiwan 114.
Organizational Affiliation: