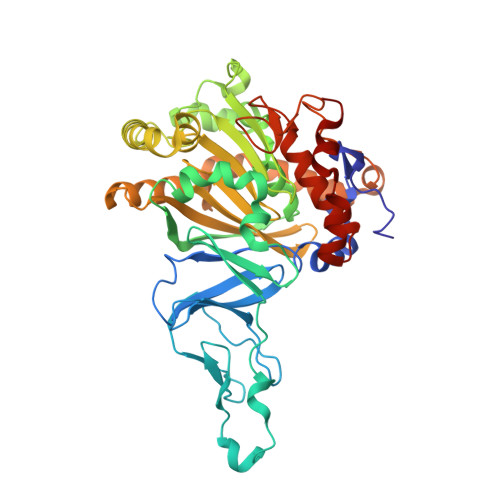
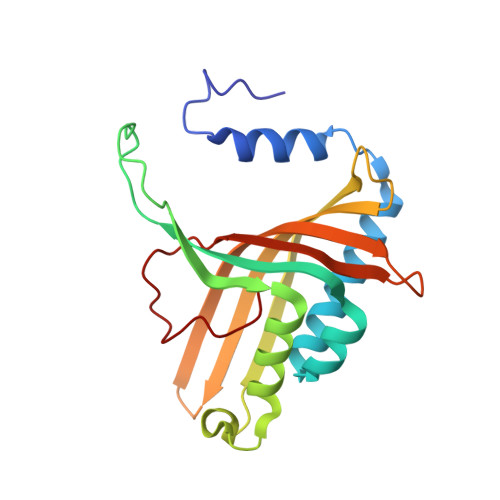
No Binding to Naphthalene Dioxygenase.
Karlsson, A., Parales, J.V., Parales, R.E., Gibson, D.T., Eklund, H., Ramaswamy, S.(2005) J Biol Inorg Chem 10: 483
- PubMed: 15942729 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-005-0657-1
- Primary Citation Related Structures:
1UUV, 1UUW - PubMed Abstract:
Nitric oxide (NO) is commonly used as an analogue for dioxygen in structural and spectroscopic studies of oxygen binding and oxygen activation. In this study, crystallographic structures of naphthalene dioxygenase (NDO) in complex with nitric oxide are reported. In the presence of the aromatic substrate indole, NO is bound end-on to the active-site mononuclear iron of NDO. The structural observations correlate well with spectroscopic measurements of NO binding to NDO in solution. However, the end-on binding of NO is in contrast to the recently reported structure of oxygen to the active-site iron of NDO that binds side-on. While NO is a good oxygen analogue with many similarities to O(2), the different binding mode of NO to the active-site iron atom leads to different mechanistic implications. Hence, caution needs to be used in extrapolating NO as an analogue to O(2) binding.
- Department of Molecular Biology, Biomedical Center, Swedish University of Agricultural Sciences, 75124 Uppsala, Sweden.
Organizational Affiliation: