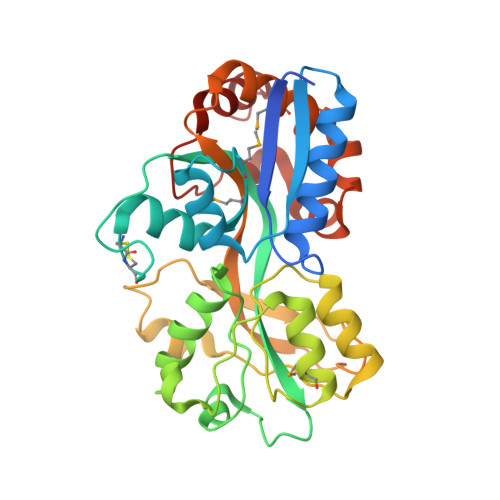
Structure of the Thermus Thermophilus Putative Periplasmic Glutamate/Glutamine-Binding Protein
Takahashi, H., Inagaki, E., Kuroishi, C., Tahirov, T.H.(2004) Acta Crystallogr D Biol Crystallogr 60: 1846
- PubMed: 15388932 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904019420
- Primary Citation Related Structures:
1US4, 1US5 - PubMed Abstract:
As part of a structural genomics project, the crystal structure of a 314-amino-acid protein encoded by Thermus thermophilus HB8 gene TT1099 was solved to 1.75 A using the multiple-wavelength anomalous dispersion (MAD) method and a selenomethionine-incorporated protein. The native protein structure was solved to 1.5 A using the molecular-replacement method. Both structures revealed a bound ligand, L-glutamate or L-glutamine, and a fold related to the periplasmic substrate-binding proteins (PSBP). Further comparative structural analysis with other PSBP-fold proteins revealed the conservation of the predicted membrane permease binding surface area and indicated that the T. thermophilus HB8 molecule is most likely to be an L-glutamate and/or an L-glutamine-binding protein related to the cluster 3 periplasmic receptors. However, the geometry of ligand binding is unique to the T. thermophilus HB8 molecule.
- Highthroughput Factory, RIKEN Harima Institute, 1-1-1 Kouto, Mikazuki-cho, Sayo-gun, Hyogo 679-5148, Japan.
Organizational Affiliation: