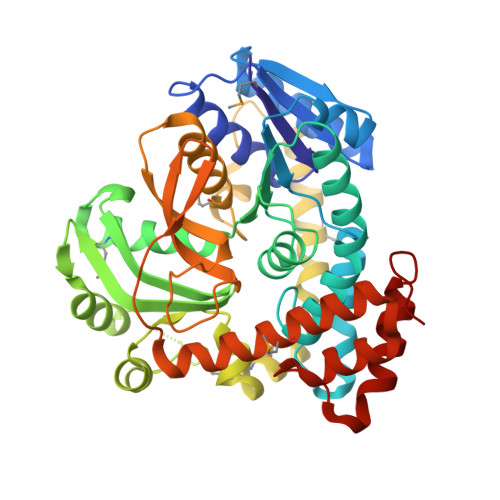
An Unusual Mechanism of Glycoside Hydrolysis Involving Redox and Elimination Steps by a Family 4 Beta-Glycosidase from Thermotoga Maritima.
Yip, V.L., Varrot, A., Davies, G.J., Rajan, S.S., Yang, X., Thompson, J., Anderson, W.F., Withers, S.G.(2004) J Am Chem Soc 126: 8354
- PubMed: 15237973 Search on PubMed
- DOI: https://doi.org/10.1021/ja047632w
- Primary Citation Related Structures:
1UP6 - PubMed Abstract:
Among the numerous well-characterized families of glycosidases, family 4 appears to be the anomaly, requiring both catalytic NAD+ and a divalent metal for activity. The unusual cofactor requirement prompted the proposal of a mechanism involving key NAD+-mediated redox steps as well as elimination of the glycosidic oxygen. Primary kinetic isotope effects for the 2- and 3-deutero substrate analogues, isotopic exchange with solvent, and structural analysis of a 6-phospho-beta-glucosidase, BglT (E.C. 3.2.1.6), provided evidence in support of the proposed mechanism, which has striking resemblances to that of the sugar dehydratases. Furthermore, analysis of the stereochemical outcome indicated that family 4 enzymes are retaining glycosidases.
- Department of Chemistry, University of British Columbia, Vancouver, BC, Canada V6T 1Z1.
Organizational Affiliation: