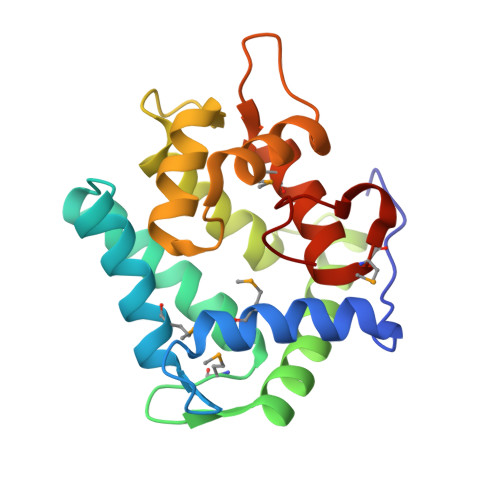
The crystal structures of semi-synthetic aequorins
Toma, S., Chong, K.T., Nakagawa, A., Teranishi, K., Inouye, S., Shimomura, O.(2005) Protein Sci 14: 409-416
- PubMed: 15632284 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.041067805
- Primary Citation Related Structures:
1UHH, 1UHI, 1UHJ, 1UHK - PubMed Abstract:
The photoprotein aequorin emits light by an intramolecular reaction in the presence of a trace amount of Ca(2+). Semi-synthetic aequorins, produced by replacing the coelenterazine moiety in aequorin with the analogues of coelenterazine, show widely different sensitivities to Ca(2+). To understand the structural basis of the Ca(2+)-sensitivity, we determined the crystal structures of four semi-synthetic aequorins (cp-, i-, br- and n-aequorins) at resolutions of 1.6-1.8 A. In general, the protein structures of these semi-synthetic aequorins are almost identical to native aequorin. Of the four EF-hand domains in the molecule, EF-hand II does not bind Ca(2+), and the loop of EF-hand IV is clearly deformed. It is most likely that the binding of Ca(2+) with EF-hands I and III triggers luminescence. Although little difference was found in the overall structures of aequorins investigated, some significant differences were found in the interactions between the substituents of coelenterazine moiety and the amino acid residues in the binding pocket. The coelenterazine moieties in i-, br-, and n-aequorins have bulky 2-substitutions, which can interfere with the conformational changes of protein structure that follow the binding of Ca(2+) to aequorin. In cp-aequorin, the cyclopentylmethyl group that substitutes for the original 8-benzyl group does not interact hydrophobically with the protein part, giving the coelenterazine moiety more conformational freedom to promote the light-emitting reaction. The differences of various semi-synthetic aequorins in Ca(2+)-sensitivity and reaction rate are explained by the capability of the involved groups and structures to undergo conformational changes in response to the Ca(2+)-binding.
- Institute for Protein Research, Osaka University, 3-2 Yamadaoka, Suita, Osaka 565-0871, Japan.
Organizational Affiliation: