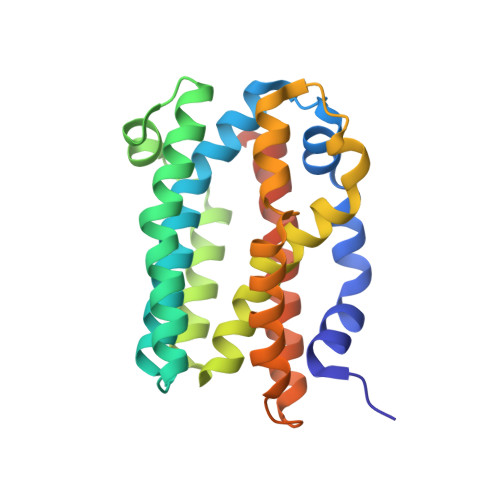
Structure analysis of PH1161 protein, a transcriptional activator TenA homologue from the hyperthermophilic archaeon Pyrococcus horikoshii.
Itou, H., Yao, M., Watanabe, N., Tanaka, I.(2004) Acta Crystallogr D Biol Crystallogr 60: 1094-1100
- PubMed: 15159569 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904008522
- Primary Citation Related Structures:
1UDD - PubMed Abstract:
The crystal structure of the Bacillus subtilis TenA-homologue protein PH1161 from the hyperthermophilic archaebacterium Pyrococcus horikoshii was determined. TenA is known to belong to a new family of activators that stimulate the production of extracellular proteases in B. subtilis. A sequence-similarity search revealed that TenA-homologue proteins are widespread in bacteria and archaea, suggesting that this family of proteins plays an essential role in these organisms. In the present study, the first three-dimensional structure of a member of the TenA family of proteins was determined, unexpectedly revealing that the protein has a fold identical to that of haem oxygenase-1. Analysis has also shown that the protein has a unique ligand-binding pocket. Electron density of a bound ligand molecule was observed in this pocket. These results provide a valuable insight into the functional understanding of the TenA family of proteins.
- Frontier Research Center for Post-genomic Science and Technology, Hokkaido University, Sapporo 001-0021, Japan.
Organizational Affiliation: