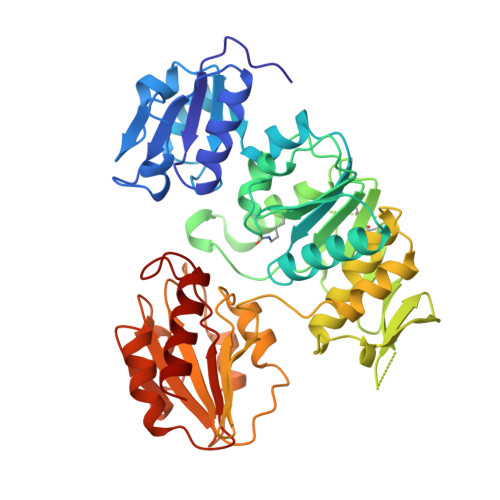
Crystal structure of UDP-N-acetylmuramoyl-L-alanine:D-glutamate ligase from Escherichia coli.
Bertrand, J.A., Auger, G., Fanchon, E., Martin, L., Blanot, D., van Heijenoort, J., Dideberg, O.(1997) EMBO J 16: 3416-3425
- PubMed: 9218784 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/16.12.3416
- Primary Citation Related Structures:
1UAG - PubMed Abstract:
UDP-N-acetylmuramoyl-L-alanine:D-glutamate ligase (MurD) is a cytoplasmic enzyme involved in the biosynthesis of peptidoglycan which catalyzes the addition of D-glutamate to the nucleotide precursor UDP-N-acetylmuramoyl-L-alanine (UMA). The crystal structure of MurD in the presence of its substrate UMA has been solved to 1.9 A resolution. Phase information was obtained from multiple anomalous dispersion using the K-shell edge of selenium in combination with multiple isomorphous replacement. The structure comprises three domains of topology each reminiscent of nucleotide-binding folds: the N- and C-terminal domains are consistent with the dinucleotide-binding fold called the Rossmann fold, and the central domain with the mononucleotide-binding fold also observed in the GTPase family. The structure reveals the binding site of the substrate UMA, and comparison with known NTP complexes allows the identification of residues interacting with ATP. The study describes the first structure of the UDP-N-acetylmuramoyl-peptide ligase family.
- Institut de Biologie Structurale Jean-Pierre Ebel (CEA-CNRS), Laboratoire de Cristallographie Macromoléculaire, Grenoble, France.
Organizational Affiliation: