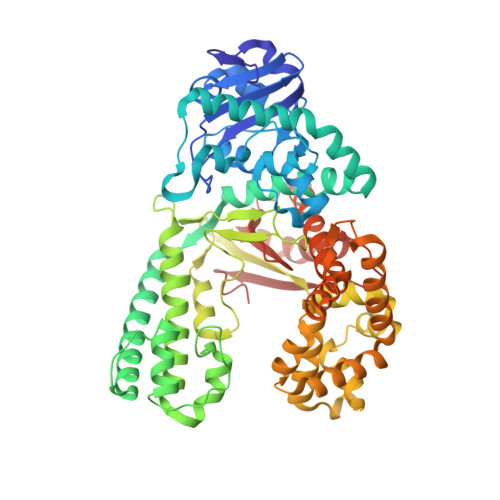
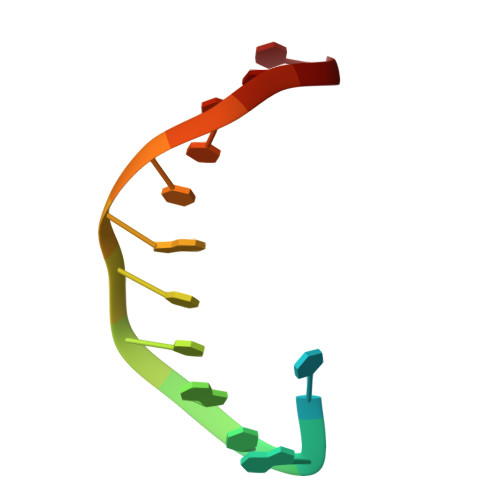
Observing translesion synthesis of an aromatic amine DNA adduct by a high-fidelity DNA polymerase
Hsu, G.W., Kiefer, J.R., Becherel, O.J., Fuchs, R.P.P., Beese, L.S.(2004) J Biological Chem 279: 50280-50285
- PubMed: 15385534 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M409224200
- Primary Citation Related Structures:
1UA0, 1UA1 - PubMed Abstract:
Aromatic amines have been studied for more than a half-century as model carcinogens representing a class of chemicals that form bulky adducts to the C8 position of guanine in DNA. Among these guanine adducts, the N-(2'-deoxyguanosin-8-yl)-aminofluorene (G-AF) and N-2-(2'-deoxyguanosin-8-yl)-acetylaminofluorene (G-AAF) derivatives are the best studied. Although G-AF and G-AAF differ by only an acetyl group, they exert different effects on DNA replication by replicative and high-fidelity DNA polymerases. Translesion synthesis of G-AF is achieved with high-fidelity polymerases, whereas replication of G-AAF requires specialized bypass polymerases. Here we have presented structures of G-AF as it undergoes one round of accurate replication by a high-fidelity DNA polymerase. Nucleotide incorporation opposite G-AF is achieved in solution and in the crystal, revealing how the polymerase accommodates and replicates past G-AF, but not G-AAF. Like an unmodified guanine, G-AF adopts a conformation that allows it to form Watson-Crick hydrogen bonds with an opposing cytosine that results in protrusion of the bulky fluorene moiety into the major groove. Although incorporation opposite G-AF is observed, the C:G-AF base pair induces distortions to the polymerase active site that slow translesion synthesis.
- Department of Biochemistry, Duke University Medical Center, Durham, North Carolina 27710, USA.
Organizational Affiliation: