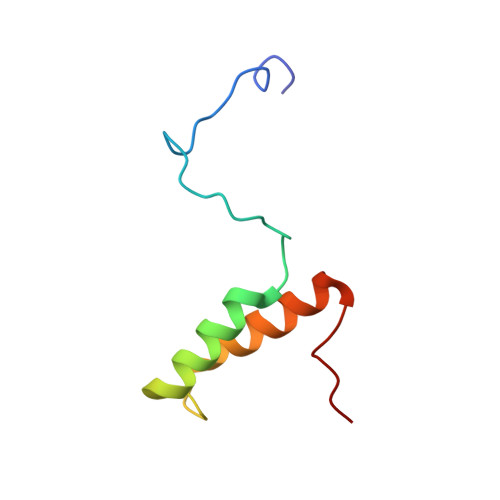
Yeast cox17 solution structure and Copper(I) binding.
Abajian, C., Yatsunyk, L.A., Ramirez, B.E., Rosenzweig, A.C.(2004) J Biological Chem 279: 53584-53592
- PubMed: 15465825 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M408099200
- Primary Citation Related Structures:
1U96, 1U97 - PubMed Abstract:
Cox17 is a 69-residue cysteine-rich, copper-binding protein that has been implicated in the delivery of copper to the Cu(A) and Cu(B) centers of cytochrome c oxidase via the copper-binding proteins Sco1 and Cox11, respectively. According to isothermal titration calorimetry experiments, fully reduced Cox17 binds one Cu(I) ion with a K(a) of (6.15 +/- 5.83) x 10(6) M(-1). The solution structures of both apo and Cu(I)-loaded Cox17 reveal two alpha helices preceded by an extensive, unstructured N-terminal region. This region is reminiscent of intrinsically unfolded proteins. The two structures are very similar overall with residues in the copper-binding region becoming more ordered in Cu(I)-loaded Cox17. Based on the NMR data, the Cu(I) ion has been modeled as two-coordinate with ligation by conserved residues Cys(23) and Cys(26). This site is similar to those observed for the Atx1 family of copper chaperones and is consistent with reported mutagenesis studies. A number of conserved, positively charged residues may interact with complementary surfaces on Sco1 and Cox11, facilitating docking and copper transfer. Taken together, these data suggest that Cox17 is not only well suited to a copper chaperone function but is specifically designed to interact with two different target proteins.
- Department of Biochemistry, Northwestern University, Evanston, IL 60208, USA.
Organizational Affiliation: