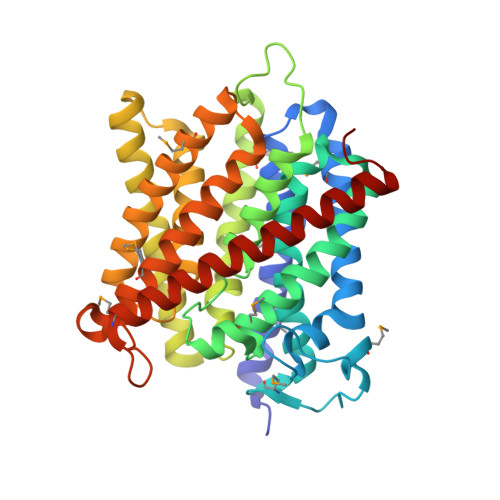
Mechanism of ammonia transport by Amt/MEP/Rh: structure of AmtB at 1.35 A
Khademi, S., O'Connell III, J., Remis, J., Robles-Colmenares, Y., Miercke, L.J.W., Stroud, R.M.(2004) Science 305: 1587-1594
- PubMed: 15361618 Search on PubMed
- DOI: https://doi.org/10.1126/science.1101952
- Primary Citation Related Structures:
1U77, 1U7C, 1U7G - PubMed Abstract:
The first structure of an ammonia channel from the Amt/MEP/Rh protein superfamily, determined to 1.35 angstrom resolution, shows it to be a channel that spans the membrane 11 times. Two structurally similar halves span the membrane with opposite polarity. Structures with and without ammonia or methyl ammonia show a vestibule that recruits NH4+/NH3, a binding site for NH4+, and a 20 angstrom-long hydrophobic channel that lowers the NH4+ pKa to below 6 and conducts NH3. Favorable interactions for NH3 are seen within the channel and use conserved histidines. Reconstitution of AmtB into vesicles shows that AmtB conducts uncharged NH3.
- Department of Biochemistry and Biophysics, S412C Genentech Hall, University of California-San Francisco, 600 16th Street, San Francisco, CA 94143-2240, USA.
Organizational Affiliation: