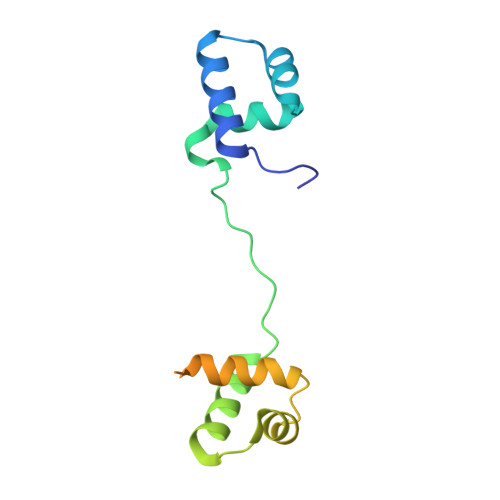
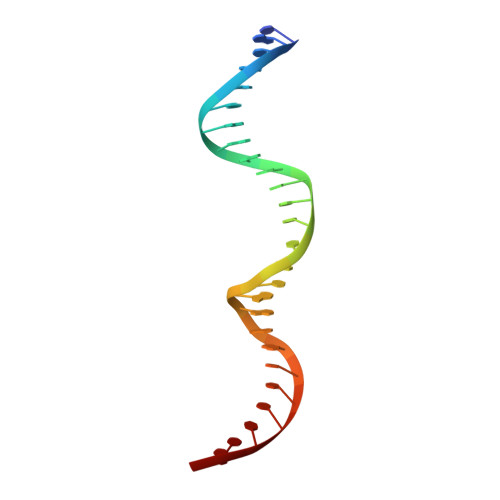
Structural analysis of the bipartite DNA-binding domain of Tc3 transposase bound to transposon DNA
Watkins, S., van Pouderoyen, G., Sixma, T.K.(2004) Nucleic Acids Res 32: 4306-4312
- PubMed: 15304566 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkh770
- Primary Citation Related Structures:
1U78 - PubMed Abstract:
The bipartite DNA-binding domain of Tc3 transposase, Tc3A, was crystallized in complex with its transposon recognition sequence. In the structure the two DNA-binding domains form structurally related helix-turn-helix (HTH) motifs. They both bind to the major groove on a single DNA oligomer, separated by a linker that interacts closely with the minor groove. The structure resembles that of the transcription factor Pax6 DNA-binding domain, but the relative orientation of the HTH-domain is different. The DNA conformation is distorted, characterized by local narrowing of the minor groove and bends at both ends. The protein-DNA recognition takes place through base and backbone contacts, as well as shape-recognition of the distortions in the DNA. Charged interactions are primarily found in the N-terminal domain and the linker indicating that these may form the initial contact area. Two independent dimer interfaces could be relevant for bringing together transposon ends and for binding to a direct repeat site in the transposon end. In contrast to the Tn5 synaptic complex, the two Tc3A DNA-binding domains bind to a single Tc3 transposon end.
- Division of Molecular Carcinogenesis, Netherlands Cancer Institute, Plesmanlaan 121, 1066 CX Amsterdam, The Netherlands.
Organizational Affiliation: