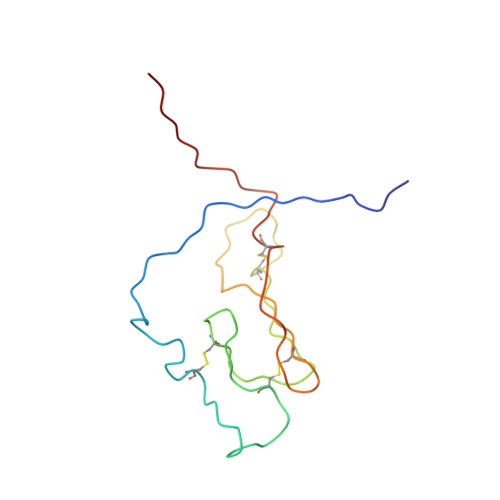
NMR structure and peptide hormone binding site of the first extracellular domain of a type B1 G protein-coupled receptor
Grace, C.R., Perrin, M.H., DiGruccio, M.R., Miller, C.L., Rivier, J.E., Vale, W.W., Riek, R.(2004) Proc Natl Acad Sci U S A 101: 12836-12841
- PubMed: 15326300 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0404702101
- Primary Citation Related Structures:
1U34 - PubMed Abstract:
The corticotropin-releasing factor (CRF) ligand family has diverse effects on the CNS, including the modulation of the stress response. The ligands' effects are mediated by binding to CRF G protein-coupled receptors. We have determined the 3D NMR structure of the N-terminal extracellular domain (ECD1) of the mouse CRF receptor 2beta, which is the major ligand recognition domain, and identified its ligand binding site by chemical-shift perturbation experiments. The fold is identified as a short consensus repeat (SCR), a common protein interaction module. Mutagenesis reveals the integrity of the hormone-binding site in the full-length receptor. This study proposes that the ECD1 captures the C-terminal segment of the ligand, whose N terminus then penetrates into the transmembrane region of the receptor to initiate signaling. Key residues of SCR in the ECD1 are conserved in the G protein-coupled receptor subfamily, suggesting the SCR fold in all of the ECD1s of this subfamily.
- Structural Biology Laboratory, and The Clayton Foundation Laboratories for Peptide Biology, The Salk Institute for Biological Studies, 10010 North Torrey Pines Road, La Jolla, CA 92037, USA. perrin@salk.edu
Organizational Affiliation: