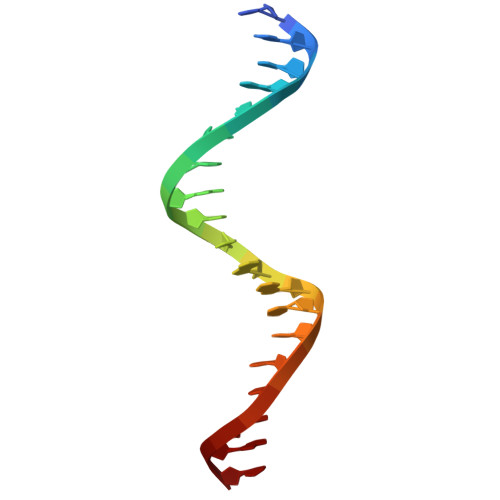
Crystal structure of trp repressor/operator complex at atomic resolution.
Otwinowski, Z., Schevitz, R.W., Zhang, R.G., Lawson, C.L., Joachimiak, A., Marmorstein, R.Q., Luisi, B.F., Sigler, P.B.(1988) Nature 335: 321-329
- PubMed: 3419502 Search on PubMed
- DOI: https://doi.org/10.1038/335321a0
- Primary Citation Related Structures:
1TRO - PubMed Abstract:
The crystal structure of the trp repressor/operator complex shows an extensive contact surface, including 24 direct and 6 solvent-mediated hydrogen bonds to the phosphate groups of the DNA. There are no direct hydrogen bonds or non-polar contacts to the bases that can explain the repressor's specificity for the operator sequence. Rather, the sequence seems to be recognized indirectly through its effects on the geometry of the phosphate backbone, which in turn permits the formation of a stable interface. Water-mediated polar contacts to the bases also appear to contribute part of the specificity.
- Department of Biochemistry and Molecular Biology, University of Chicago, Illinois 60637.
Organizational Affiliation: