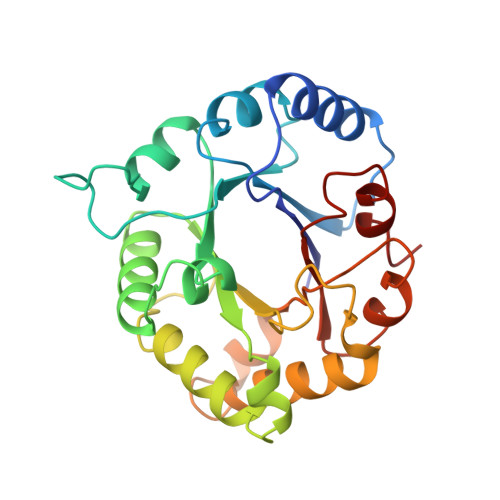
Crystal structure of recombinant chicken triosephosphate isomerase-phosphoglycolohydroxamate complex at 1.8-A resolution.
Zhang, Z., Sugio, S., Komives, E.A., Liu, K.D., Knowles, J.R., Petsko, G.A., Ringe, D.(1994) Biochemistry 33: 2830-2837
- PubMed: 8130195 Search on PubMed
- DOI: https://doi.org/10.1021/bi00176a012
- Primary Citation Related Structures:
1TPH - PubMed Abstract:
The crystal structure of recombinant chicken triosephosphate isomerase (TIM, E.C. 5.3.1.1) complexed with the intermediate analogue phosphoglycolohydroxamate (PGH) has been solved by the method of molecular replacement and refined to an R-factor of 18.5% at 1.8-A resolution. The structure is essentially identical to that of the yeast TIM-PGH complex [Davenport, R. C., et al. (1991) Biochemistry 30, 5821-5826] determined earlier and refined at comparable resolution. This identity extends to the high-energy conformations of the active-site residues Lys13 and Ser211, as well as the positions of several bound water molecules that are retained in the active site when PGH is bound. Comparison with the structure of uncomplexed chicken TIM shows that the catalytic base, Glu165, moves several angstroms when PGH binds. This movement may provide a trigger for a larger conformational change, one of 7 A, in a loop near the active site, which folds down like a lid to shield the bound inhibitor and catalytic residues from contact with bulk solvent. These same conformational changes were seen in crystalline yeast TIM upon binding of PGH; their occurrence here in a different crystal form of TIM eliminates the possibility that they are an artifact of crystal packing.
- Department of Biochemistry, Rosenstiel Basic Medical Sciences Research Center, Brandeis University, Waltham, Massachusetts 02254-9110.
Organizational Affiliation: