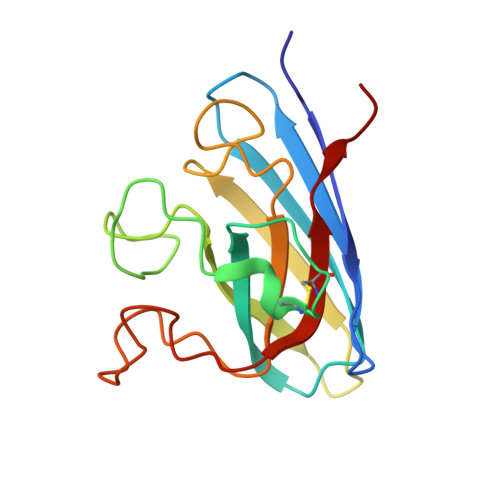
Structure of the cytosolic Cu,Zn superoxide dismutase from Schistosoma mansoni.
Cardoso, R.M., Silva, C.H., Ulian de Araujo, A.P., Tanaka, T., Tanaka, M., Garratt, R.C.(2004) Acta Crystallogr D Biol Crystallogr 60: 1569-1578
- PubMed: 15333927 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904016798
- Primary Citation Related Structures:
1TO4, 1TO5 - PubMed Abstract:
Cu,Zn superoxide dismutase (Cu,Zn SOD) is an essential enzyme for protecting cells from the toxic effects of reactive oxygen species. In humans, two distinct Cu,Zn SOD genes are located on chromosomes 4 and 21 and mutations in the latter have been associated with familial amyotrophic lateral sclerosis. Similarly, schistosomes (trematode parasites responsible for the chronically debilitating disease schistosomiasis) also produce two distinct Cu,Zn SODs, in this case one cytosolic and one bearing a signal peptide. The crystal structure of the cytosolic form of the enzyme from the human trematode Schistosoma mansoni (SmCtSOD) was solved and refined to a resolution of 2.2 A (space group P2(1)2(1)2(1), R = 17.6% and R(free) = 24.1%) and 1.55 A (space group P2(1), R = 15.7% and R(free) = 17.1%). This is the first report of a crystal structure of a Cu,Zn superoxide dismutase derived from a human parasite. Alternate positions for the catalytic copper and its water ligand were refined for the 1.55 A SmCtSOD model, but the most interesting structural differences between SmCtSOD and the human homologue reside in the loops used for electrostatic guidance of the substrate to the enzyme active site.
- Instituto de Física de São Carlos, Universidade de São Paulo, Av. Trabalhador Sãocarlense 400, São Carlos, SP, 13566-590, Brazil.
Organizational Affiliation: