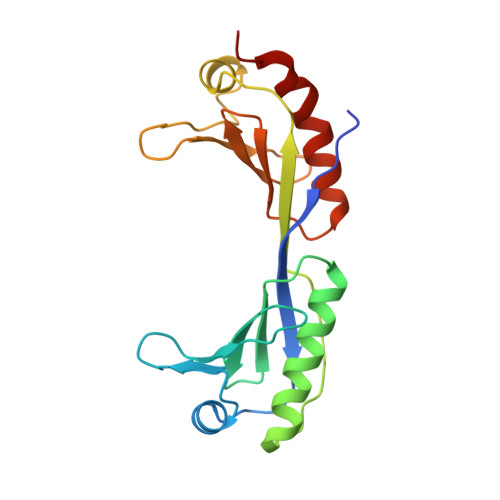
How proteins recognize the TATA box.
Juo, Z.S., Chiu, T.K., Leiberman, P.M., Baikalov, I., Berk, A.J., Dickerson, R.E.(1996) J Mol Biology 261: 239-254
- PubMed: 8757291 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1996.0456
- Primary Citation Related Structures:
1TGH - PubMed Abstract:
The crystal structure of a complex of human TATA-binding protein with TATA-sequence DNA has been solved, complementing earlier TBP/DNA analyses from Saccharomyces cerevisiae and Arabidopsis thaliana. Special insight into TATA box specificity is provided by considering the TBP/DNA complex, not as a protein molecule with bound DNA, but as a DNA duplex with a particularly large minor groove ligand. This point of view provides explanations for: (1) why T.A base-pairs are required rather than C.G; (2) why an alternation of T and A bases is needed; (3) how TBP recognizes the upstream and downstream ends of the TATA box in order to bind properly; and (4) why the second half of the TATA box can be more variable than the first.
- Molecular Biology Institute, University of California at Los Angeles 90095, USA.
Organizational Affiliation: