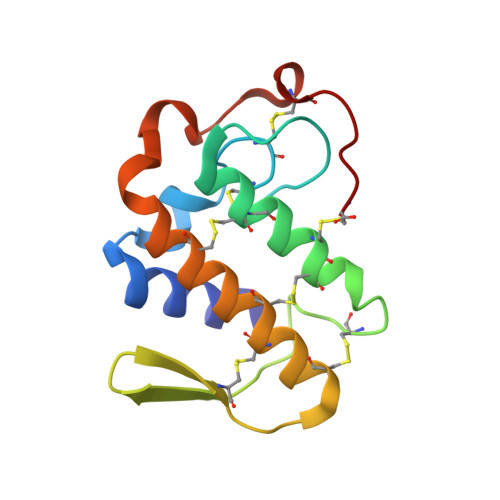
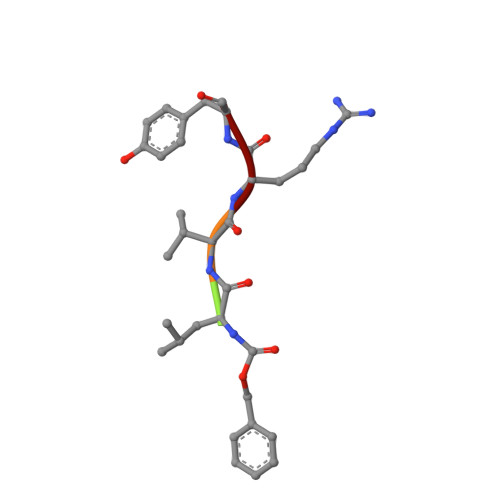
Crystal Structure of the complex formed between russells viper phospholipase A2 and a designed peptide inhibitor Cbz-dehydro-Leu-Val-Arg-Tyr at 1.2A resolution
Singh, N., Kaur, P., Somvanshi, R.K., Sharma, S., Dey, S., Perbandt, M., Betzel, C., Singh, T.P.To be published.